Measuring covariate shift
In this vignette, we will measure the reconstruction error of a PCA, in order to measure the amount of covariate shift between the training data and the data used for prediction.
using SpeciesDistributionToolkit
const SDT = SpeciesDistributionToolkit
using CairoMakieWe will need to rely on the integration with MultivariateStats in order to fit and then apply the PCA, so we will load this package as well:
using Statistics
using MultivariateStats This is a temporary vignette!
This functionality will ultimately be ported to the SDeMo package. It is included as a reference, but the code will be updated to reflect the new functions.
Declaring the model
We will measure the multi-variate covariate shift, which reflects how much the joint distribution of variables shifts from the training data to the prediction data. In order to do this, we first need a model:
model = SDM(PCATransform, Logistic, SDeMo.__demodata()...)
variables!(model, ForwardSelection)☑️ PCATransform → Logistic → P(x) ≥ 0.575 🗺️We then collect the data from the CHELSA2 BioClim on which this model was trained, as a series of layers:
pol = getpolygon(PolygonData(OpenStreetMap, Places); place = "Corse")
L = SDMLayer{Float32}[
SDMLayer(RasterData(CHELSA2, BioClim); SDT.boundingbox(pol)..., layer = i) for
i in 1:19
]
mask!(L, pol)19-element Vector{SDMLayer{Float32}}:
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)
🗺️ A 205 × 124 layer (13685 Float32 cells)Reconciling a stack of layers
In rare cases, layers from the same providers may have minute differences in their x and y coordinates after being clipped. In order to repair them, we can use the reconcile function or its mutating variant. This should only be used when the layers have non-matching coordinates.
reconcile!(L)PCA on the training data
Our first step is to fit a PCA on the training data. Because we are working with a dataset with variables on different scales, we will transform all the variables so that they have a mean of 0 and unit variance.
X = features(model)[variables(model),:]
μ = vec(mean(X, dims=2))
σ = vec(mean(X, dims=2))
X = (X .- μ) ./ σ;We can now fit a PCA on this matrix. The package supports fitting directly from layers as well.
M = fit(PCA, X)PCA(indim = 4, outdim = 1, principalratio = 0.9912894019942406)
Pattern matrix (unstandardized loadings):
────────────
PC1
────────────
1 2.76728
2 0.032147
3 -0.188889
4 -0.141518
────────────
Importance of components:
───────────────────────────────────
PC1
───────────────────────────────────
SS Loadings (Eigenvalues) 7.71455
Variance explained 0.991289
Cumulative variance 0.991289
Proportion explained 1.0
Cumulative proportion 1.0
───────────────────────────────────Measuring the reconstruction error
We now need to transform the layers, but only after ensuring that they are transformed using the same operations that gave us a training dataset with mean of 0 and unit variance.
K = (L[variables(model)] .- μ)./σ4-element Vector{SDMLayer{Float64}}:
🗺️ A 205 × 124 layer (13685 Float64 cells)
🗺️ A 205 × 124 layer (13685 Float64 cells)
🗺️ A 205 × 124 layer (13685 Float64 cells)
🗺️ A 205 × 124 layer (13685 Float64 cells)To measure the covariate shift, we are going to take the layers, project them using the PCA we trained on the model training data; we will then attempt to reconstruct the original data from their projection:
Y = reconstruct(M, predict(M, K));Note that this is returned as a series of layers.
In order to measure the difference between the original layers and their reconstruction, we will take the Frobenius norm of the difference between the two matrices.
N = SpeciesDistributionToolkit._X_from_layers(K);We will now get the total reconstruction error for this matrix. Note that we are converting the vector of layers into a Matrix here:
P = Matrix(Y .- K)
import LinearAlgebra
LinearAlgebra.norm(P)33.381825033814394Mapping the reconstruction error
We will now look at the distance between the actual value and the reconstructed value, for each cell in the layer.
D = mosaic(x -> sqrt(sum(x.^2)), (Y .- K))🗺️ A 205 × 124 layer with 13685 Float64 cells
Spatial Reference System: +proj=longlat +datum=WGS84 +no_defsWhen should layers become matrices?
Never? A lot of operations that would work on matrices will also work on layers. Our suggestion is to only rely on the conversion to matrices when required to interact with other packages. When the layers are converted to a matrix, the reconcile function will be applied if necessary.
This layer measures the per-pixel error in the reconstruction, based on a PCA fitted on the training data. It can be used to map the areas where the extant predictions are further away from the training data.
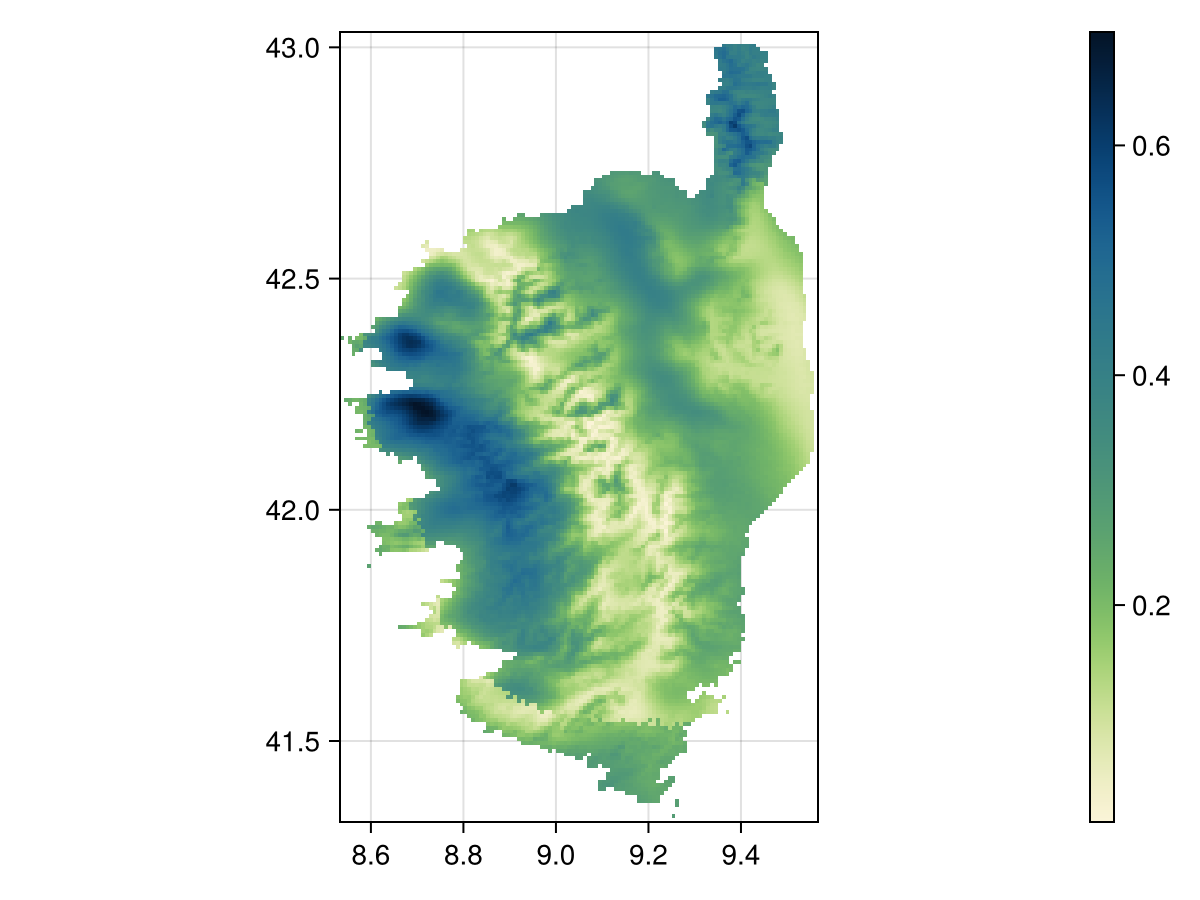
Code for the figure
f = Figure()
ax = Axis(f[1,1]; aspect=DataAspect())
hm = heatmap!(ax, D, colormap=Reverse(:navia))
Colorbar(f[1,2], hm)Related documentation
SimpleSDMLayers.reconcile! Function
reconcile!(L::Vector{<:SDMLayer})Takes a stack of layers, and ensure that they have the same x and y values. This should be used as a last resort when some data providers have layers that don't fully align after clipping.
SimpleSDMLayers.reconcile Function
reconcile(L::Vector{<:SDMLayer})Non-mutating version of reconcile!.