Feature importance
The purpose of this vignette is to show how to measure the importance of features in a model.
using SpeciesDistributionToolkit
using Statistics
using PrettyTables
using CairoMakieWe will train a very simple model using the demonstration dataset:
model = SDM(RawData, Logistic, SDeMo.__demodata()...)❎ RawData → Logistic → P(x) ≥ 0.5 🗺️We will then select variables using forward selection, but retaining the 1st and 12th variables (mean temperature, total precipitation).
variables!(model, ForwardSelection; included = [1, 12])
variables(model)8-element Vector{Int64}:
1
12
18
4
2
14
13
11Variable selection
For more on variable selection, see the vignette.There are additional techniques to decide which variables are included in a model.
We will use three techniques for feature importance: permutation importance, partial dependence, and Shapley importance.
Permutation importance
We will split the instances into a training and validation group using holdout – this will be useful to illustrate the permutation importance, which works best when applied to a dataset that was not used for model training.
train, test = holdout(model);Splitting the dataset is important here, because there is a risk of wrongly evaluating the importance of features if the model overfits. The other two methods can be applied on a trained model without splitting.
There are several ways to perform this task, the first being by passing an array with the positions of training/testing instances:
featureimportance(PermutationImportance, model, 1, train, test)0.8276759044187986Interpretation of importance
The ratio is measured as permuted score over true score, and the difference as permuted score minus true score. For this reason, the interpretation must depend on whether a higher value of the measure is associated to a better performance of the model. For cross-entropy loss, lower values are best.
The interpretation of this value depends on a few things, which are all set through keyword arguments. The first is the optimality measure – by default, this is the mcc. The second is whether is the importance is measured as a ratio (the default, true) or a difference. For example, we can measure the importance of the same variable as "how much it decreases the F₁ measure when the data are shuffled" with
featureimportance(
PermutationImportance,
model,
1,
train,
test;
optimality = f1,
ratio = false,
)-0.08012369314431411We can also ask the importance for the model score (as opposed to the binary prediction):
featureimportance(
PermutationImportance,
model,
1,
train,
test;
optimality = crossentropyloss,
threshold = false,
)1.4974772836047998Because this value is larger than unity, this means that the loss when this feature is shuffled is larger than when the true values are used: it contributes positively to model performance.
Note that we can also get the feature importance on all the training data:
featureimportance(PermutationImportance, model, 1; optimality = mcc)0.8229508426325901The feature importances are relative values, so it is a good idea to express them as a proportion of all the variables used in the model:
permfi = [1 - featureimportance(PermutationImportance, model, v; ratio=false, threshold=false) for v in variables(model)]
permfi ./= sum(permfi)8-element Vector{Float64}:
0.11659467910145288
0.1328074202726076
0.11448010160415181
0.1446590066612174
0.1232460941079025
0.11646780309058696
0.11850185829227323
0.13324303686980754Partial dependence importance
We can also derive a measure of feature importance from the partial dependence plot of the model; this is covered in more depth in the vignette on interpretability.
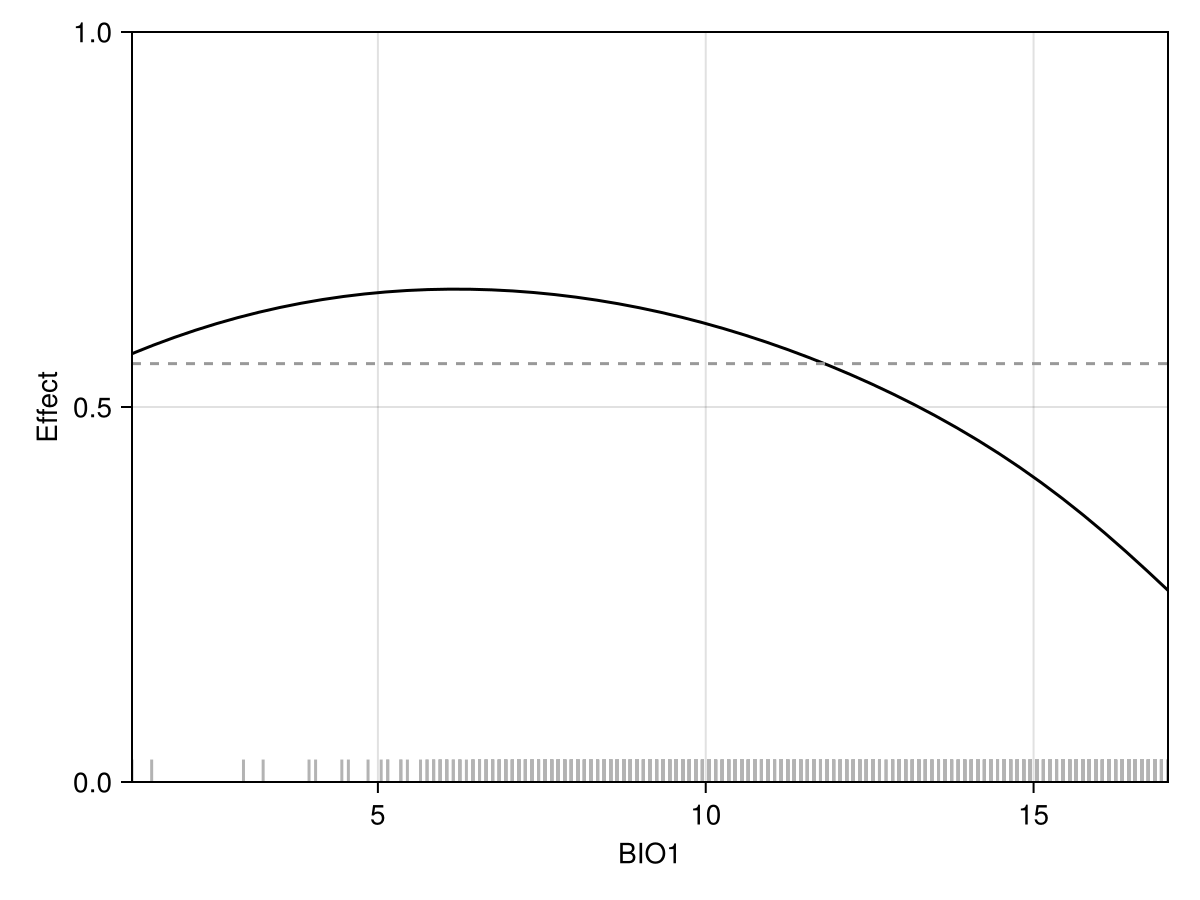
Code for the figure
f = Figure()
ax = Axis(f[1,1], xlabel="BIO1", ylabel="Effect")
vlines!(ax, features(model, 1), ymax=0.03, color=:grey70)
x, y = explainmodel(PartialDependence, model, 1, 50; threshold=false)
lines!(ax, x, y, color=:black)
hlines!(ax, mean(y), linestyle=:dash, color=:grey60)
tightlimits!(ax)
ylims!(ax, 0, 1)Essentially, the more each point is far away from the average of the partial dependence curve (indicated by a dashed line on the above figure), the more this feature is important.
Unlike with the permutation approach, this measure of feature importance does not take additional arguments. It can be applied without model re-training, as it measures importance on the model response.
partfi = [
featureimportance(PartialDependence, model, v; threshold = false) for
v in variables(model)
]
partfi ./= sum(partfi)8-element Vector{Float64}:
0.10996818330693602
0.1583874753592883
0.13453718851486607
0.20283635085953497
0.07537010249535132
0.06390023543263584
0.07032032167294011
0.1846801423584472Shapley values importance
The ShapleyMC explanation can also be used to evaluate the importance of a feature:
featureimportance(ShapleyMC, model, 1; threshold=false)0.08241609896158368This approach to feature importance is useful because it informs on the contribution of each feature to moving the prediction away from the average prediction.
shapfi = [
featureimportance(ShapleyMC, model, v; threshold = false) for
v in variables(model)
]
shapfi ./= sum(shapfi)8-element Vector{Float64}:
0.11128203042560812
0.1595638618869548
0.09329583741215314
0.1566662472266634
0.14980926145882617
0.08096500230179238
0.10254113982283986
0.145876619465162Comparison of importance values
We can finally look at the relative importance of the variables, sorted here by their importance according to the partial dependence curve:
M = hcat("BIO" .* string.(variables(model)), permfi, partfi, shapfi)
pretty_table(
M[sortperm(M[:,3]; rev=true),:],
alignment = [:l, :c, :c, :c],
backend = :markdown,
column_labels = ["Variable", "Permut.", "Part. dep.", "Shapley"],
formatters = [fmt__printf("%4.3f", [2, 3, 4])],
)| Variable | Permut. | Part. dep. | Shapley |
|---|---|---|---|
| BIO4 | 0.145 | 0.203 | 0.157 |
| BIO11 | 0.133 | 0.185 | 0.146 |
| BIO12 | 0.133 | 0.158 | 0.160 |
| BIO18 | 0.114 | 0.135 | 0.093 |
| BIO1 | 0.117 | 0.110 | 0.111 |
| BIO2 | 0.123 | 0.075 | 0.150 |
| BIO13 | 0.119 | 0.070 | 0.103 |
| BIO14 | 0.116 | 0.064 | 0.081 |
Related documentation
SDeMo.featureimportance Function
featureimportance(PermutationImportance, model, variable; kwargs...)Returns the feature importance of a variable for a model over the entire model training data. Note that to avoid overfitting, it is better to use a different testing set to evaluate the importance of features.
featureimportance(PermutationImportance, model, variable, X, y; kwargs...)Returns the feature importance of a variable for a model by predicting over features X with a ground truth vector y .
The samples keyword (defaults to 50) is the number of samples to use to evaluate the average importance.
When the ratio keyword is true, the importance is evaluated as the permuted performance divided by the original performance (values lower than 1 indicate that the variable is increasingly important). When it is false, the performance is measured as a difference (values above 0 indicate that the variable is increasingly important).
The optimality keyword takes a function (defaults to mcc) that operates on a a vector of predictions and a vector of correct labels. The interpretation of feature importance assumes that higher values of this measure correspond to a better model.
The other kwargs... are passed to predict. For example, threshold=false, optimality=crossentropyloss is the way to measure the importance of variables based on their effect on cross-entropy loss.
featureimportance(PartialDependence, model, variable; kwargs...)Measures the feature importance of a variable by measuring the distance of all points in the partial dependence curve to all other points in the curve, and then averaging it. Low values indicates a flat PDP (feature is less important). This method can be applied on a fully trained model as it focuses on the response, not the individual predictions.
featureimportance(PermutationImportance, model, variable, training, testing; kwargs...)Calculates the feature importance of a model by re-training a copy of the model using the training instances, then evaluating the feature importance on the testing instances.
Other kwargs... are passed to featureimportance.
featureimportance(ShapleyMC, model, variable; kwargs...)Returns the importance of a feature according to Shapley values; this is the average of the absolute values of this feature on the average prediction (and therefore, this is conceptually very close to the PartialDependence measure of feature importance).
Other kwargs... are passed to predict.
featureimportance(ShapleyMC, model, variable, X::Matrix; kwargs...)Returns the importance of a feature according to Shapley values; this is the average of the absolute values of this feature on the average prediction (and therefore, this is conceptually very close to the PartialDependence measure of feature importance). The Shapley values are measured on the matrix of instances X.
Other kwargs... are passed to predict.