Producing the PR and ROC curves
The purpose of this vignette is to show how to trace the PR and ROC curves from SDeMo models.
using SpeciesDistributionToolkit
using CairoMakie
using PrettyTablesWe will work on the demo data:
sdm = SDM(PCATransform, DecisionTree, SDeMo.__demodata()...)❎ PCATransform → DecisionTree → P(x) ≥ 0.5 🗺️We can now set a series of points at which to evaluate the threshold. Because we are calling the cross-validation function, this will not retrain our model, and therefore it is safe to apply this method on an already trained model.
thresholds = LinRange(0.01, 0.99, 50)
cv = [crossvalidate(sdm, montecarlo(sdm; n = 5); thr = t) for t in thresholds];We can then plot the output:
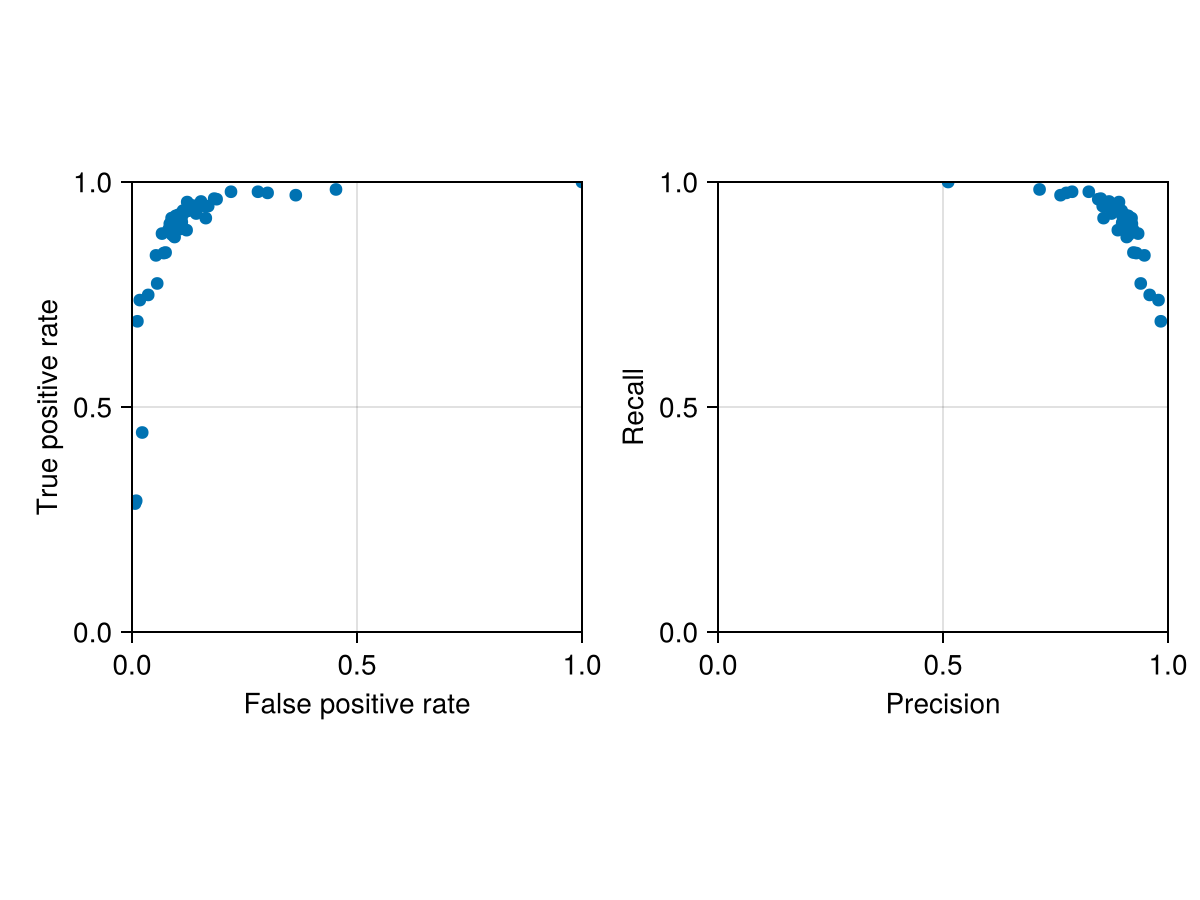
Code for the figure
f = Figure()
ax =
Axis(f[1, 1]; aspect = 1, xlabel = "False positive rate", ylabel = "True positive rate")
scatter!(ax, [fpr(s.validation) for s in cv], [tpr(s.validation) for s in cv])
ax2 = Axis(f[1, 2]; aspect = 1, xlabel = "Precision", ylabel = "Recall")
scatter!(
ax2,
[SDeMo.precision(s.validation) for s in cv],
[SDeMo.recall(s.validation) for s in cv],
)
xlims!(ax, 0.0, 1.0)
ylims!(ax, 0.0, 1.0)
xlims!(ax2, 0.0, 1.0)
ylims!(ax2, 0.0, 1.0)We can then train the model:
train!(sdm)☑️ PCATransform → DecisionTree → P(x) ≥ 0.172 🗺️It is a good idea to also perform a sanity check, by verifying that the distribution of predicted values makes sense knowing their true label (the threshold is in red):
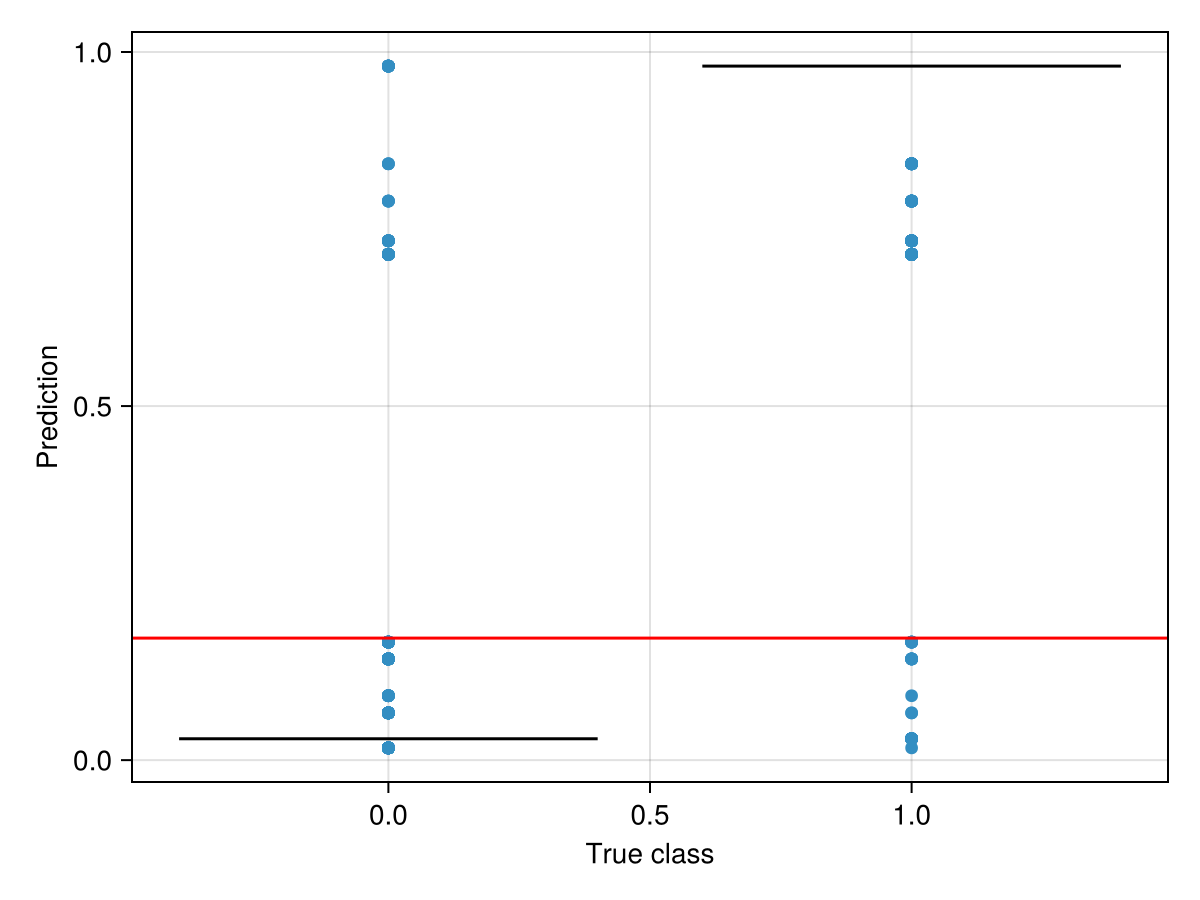
Code for the figure
f = Figure()
ax = Axis(f[1, 1]; xlabel = "True class", ylabel = "Prediction")
boxplot!(ax, labels(sdm), predict(sdm; threshold = false))
hlines!(ax, [threshold(sdm)]; color = :red)Related documentation
SDeMo.crossvalidate Function
crossvalidate(sdm, folds; thr = nothing, kwargs...)Performs cross-validation on a model, given a vector of tuples representing the data splits. The threshold can be fixed through the thr keyword arguments. All other keywords are passed to the train! method.
This method returns two vectors of ConfusionMatrix, with the confusion matrix for each set of validation data first, and the confusion matrix for the training data second.
crossvalidate(sdm::T, args...; kwargs...) where {T <: AbstractSDM}Performs cross-validation using 10-fold validation as a default. Called when crossvalidate is used without a folds second argument.