Generating neutral landscapes
The purpose of this vignette is to demonstrate how we can use the NeutralLandscapes package to generate random spatial structures.
This functionality is supported through an extension, which is only active when the NeutralLandscapes package is loaded.
using SpeciesDistributionToolkit
using CairoMakie
using NeutralLandscapes Precompiling packages...
Info Given MakieExtension was explicitly requested, output will be shown live [0K
[0KWARNING: Method definition plot!(Makie.Plot{Phylopic.silhouetteplot, var"#s38"} where var"#s38"<:Tuple{AbstractArray{var"#s37", 1} where var"#s37"<:Real, AbstractArray{var"#s36", 1} where var"#s36"<:Real, AbstractArray{var"#s35", 1} where var"#s35"<:Phylopic.PhylopicSilhouette}) in module MakieExtension at /home/runner/work/SpeciesDistributionToolkit.jl/SpeciesDistributionToolkit.jl/Phylopic/ext/MakieExtension.jl:45 overwritten at /home/runner/work/SpeciesDistributionToolkit.jl/SpeciesDistributionToolkit.jl/Phylopic/ext/MakieExtension.jl:55.
[0KERROR: Method overwriting is not permitted during Module precompilation. Use `__precompile__(false)` to opt-out of precompilation.
5527.9 ms ? Phylopic → MakieExtension
WARNING: Method definition plot!(Makie.Plot{Phylopic.silhouetteplot, var"#s38"} where var"#s38"<:Tuple{AbstractArray{var"#s37", 1} where var"#s37"<:Real, AbstractArray{var"#s36", 1} where var"#s36"<:Real, AbstractArray{var"#s35", 1} where var"#s35"<:Phylopic.PhylopicSilhouette}) in module MakieExtension at /home/runner/work/SpeciesDistributionToolkit.jl/SpeciesDistributionToolkit.jl/Phylopic/ext/MakieExtension.jl:45 overwritten at /home/runner/work/SpeciesDistributionToolkit.jl/SpeciesDistributionToolkit.jl/Phylopic/ext/MakieExtension.jl:55.
ERROR: Method overwriting is not permitted during Module precompilation. Use `__precompile__(false)` to opt-out of precompilation.This is simply achieved through an overload of the SDMLayer constructor, taking a neutral landscape maker, and a size:
L = SDMLayer(DiamondSquare(), (100, 100))🗺️ A 100 × 100 layer with 10000 Float64 cells
Spatial Reference System: +proj=longlat +datum=WGS84 +no_defs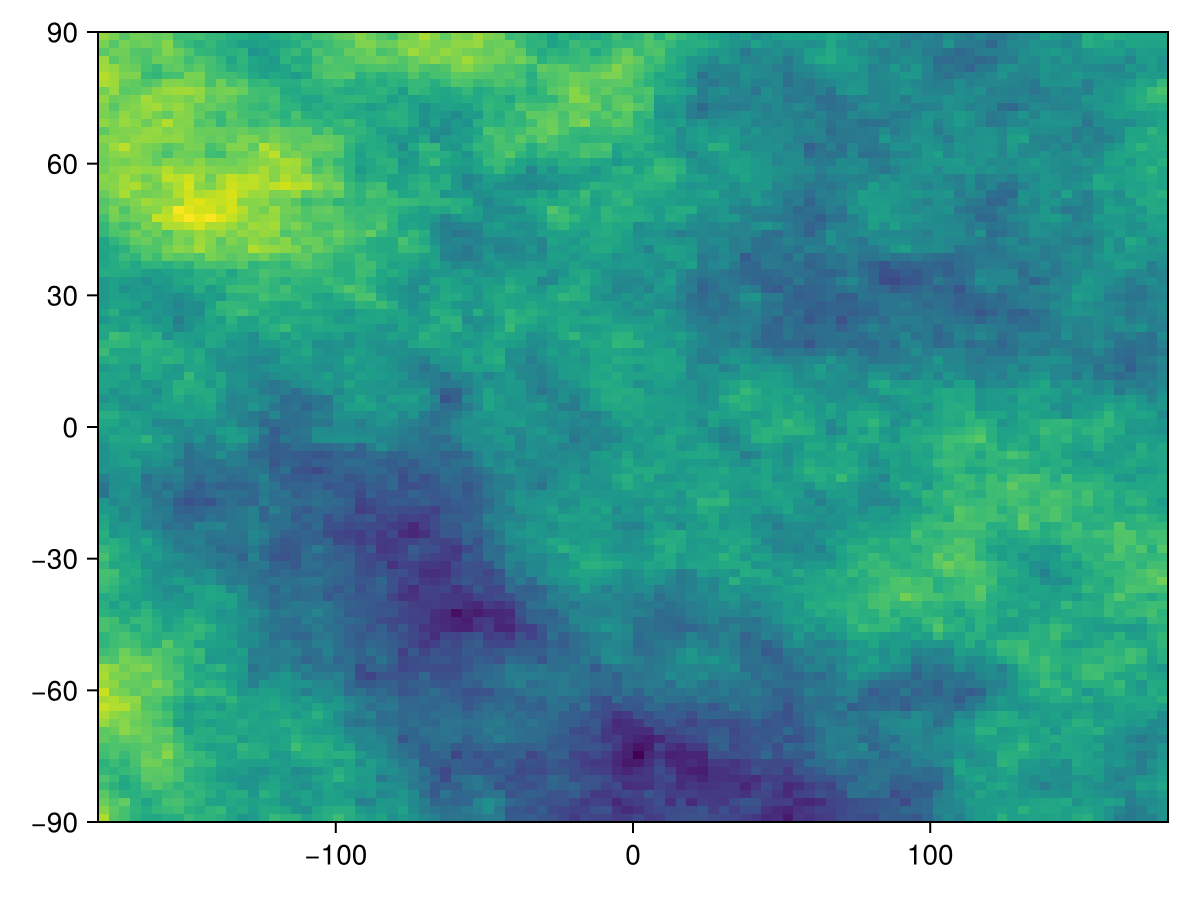
Code for the figure
heatmap(L)If we have a layer already, it can be used in place of the size argument, in which case the mask of the layer is also preserved. This is useful to rapidly fill layers that have the same structure with random data.
Finally, the SDMLayer constructor, when used this way, accepts keyword arguments like the crs of the layer, as well as its x and y fields. This is important if you want to ensure that the generated data can be overlaid on an actual layer with coordinates and projections.
Note that by default, the returned layer will be entirely filled. You can mask parts of it using nodata, or by masking it with a layer containing actual data.
Related documentation
SimpleSDMLayers.nodata Function
nodata(layer::SDMLayer, args...)Makes a copy and calls nodata! on it
SimpleSDMLayers.nodata! Function
nodata!(layer::SDMLayer{T}, nodata::T) where {T}Changes the value of the layer representing no data. This modifies the layer passed as its first argument.
sourcenodata!(layer::SDMLayer{T}, f)Removes the data matching a function
sourcenodata!(layer::SDMLayer, v)Turns off all cells containing the value v. This is an overload only applied when v is not of the correct type for layer.
SimpleSDMLayers.mask! Function
mask!(layer::SDMLayer, template::SDMLayer)Updates the positions in the first layer to be those that have a value in the second layer.
sourceSimpleSDMLayers.mask!(layer::SDMLayer, poly::T) where T<:Union{Polygon,MultiPolygon}Turns off all the cells outside the polygon (or within holes in the polygon). This modifies the object.
source