Getting started
In this tutorial, we will work on a dataset of observations of the Ring Ouzel (Turdus torquatus) in Switzerland. The purpose of the tutorial is to introduce most of the concepts that are important when working with SpeciesDistributionToolkit, by conducting an analysis from start to finish. We are very much going for breadth, not depth; in particular, this tutorial does not cover any of the theory about species distributions or species distribution models.
At many points throughout the tutorial, there will be links to other vignettes in this manual. They are not meant to be read immediately when they are mentionned, but rather consulted later on to get access to more options and choices for the analyses that are presented here.
using SpeciesDistributionToolkit
using Statistics
using PrettyTablesAbout this tutorial
This tutorial is inspired by the the excellent introduction to SDMs by Damaris Zurell. It uses the same species, but a slightly different dataset.
Most of the data types in SpeciesDistributionToolkit can be used with Makie for plotting. This is true for layers, polygons, and more. These are all documented in the "Manual" section of the documentation, under "Data visualization". To benefit from this integration, we will load the CairoMakie backend.
using CairoMakieAs in the rest of the manual, the code to produce the figures is hidden at first. This is because figure code is very verbose, and not specific to the package. Feel free to reveal the code for the figure if you want to see exactly how they are produced.
Getting the data
We start by downloading a polygon that will be used to delineate our area of interest. For this tutorial, this will be the country of Szwitzerland.
aoi = getpolygon(PolygonData(NaturalEarth, Countries); resolution = 10)["Switzerland"]Polygon Feature
├ Region => Europe
├ Subregion => Western Europe
├ Name => Switzerland
└ Population => 8574832There are several polygon providers that come built into the package, and they can be accessed via the "Datasets" menu in the navigation bar at the top of your screen. Some of these polyon providers are about administrative boundaries (GADM, Natural Earth, ESRI), but some others are about eco/bioregions.
To ensure that we only load the raster data that are within the range of our problem, we will first calculate the spatial extent of the polygon we just downloaded. The boundingbox function will work on any object that has spatial coordinates (occurrences, models, layers, etc), and return the bounding box in longitude/latitude.
const SDT = SpeciesDistributionToolkit
extent = SDT.boundingbox(aoi)(left = 5.954809188842773, right = 10.46662712097168, bottom = 45.820716857910156, top = 47.80116653442383)Alias for packages
Whenever the names of functions in SpeciesDistributionToolkit are not distinctive enough, the functions are not exported. In order to simplify the syntax, declaring a const with a shortened name (as we did just before) is a valid solution, and also introduces no performance cost. This will happen in almost all vignettes.
We will now download the entire suite of 19 BIOCLIM data from the CHELSA2 database. Before downloading, we will look at the first five layers.
chelsa_bioclim = RasterData(CHELSA2, BioClim)
layers(chelsa_bioclim)[1:5]5-element Vector{String}:
"BIO1"
"BIO2"
"BIO3"
"BIO4"
"BIO5"We can also get their full-text descriptions:
[
layerdescriptions(chelsa_bioclim)[l]
for l in layers(chelsa_bioclim)[1:5]
]5-element Vector{String}:
"Annual Mean Temperature"
"Mean Diurnal Range (Mean of monthly (max temp - min temp))"
"Isothermality (BIO2/BIO7) (×100)"
"Temperature Seasonality (standard deviation ×100)"
"Max Temperature of Warmest Month"We have enough information to download these layers, and load them in memory. It is important, whenever possible, to give a spatial extent (boundingbox) to the function to read layers. This avoids loading pretty voluminous data into memory when it is not strictly needed.
Data are saved locally
Both polygons and raster data are saved locally, which means that the first download takes longer, but subsequent reads are much faster.
The SDMLayer function is the main constructor for layers (and can also read from STAC catalogues and local files). When given a RasterProvider, it will download (or open) the relevant files. Note that we are passing the area of interest (a polygon) as the second argument, so that the raster will only be returned within the boundingbox of this geometry.
L = SDMLayer{Float32}[
SDMLayer(
chelsa_bioclim,
aoi;
layer = v
) for v in layers(chelsa_bioclim)
];Accessing raster data
For more information about accessing raster data, refer to the vignette. All datasets (and the complete set of keyword options) are also listed in the navigation bar at the top of the screen.
When these layers are first downloaded, they will cover the entire extent of the bounding box:
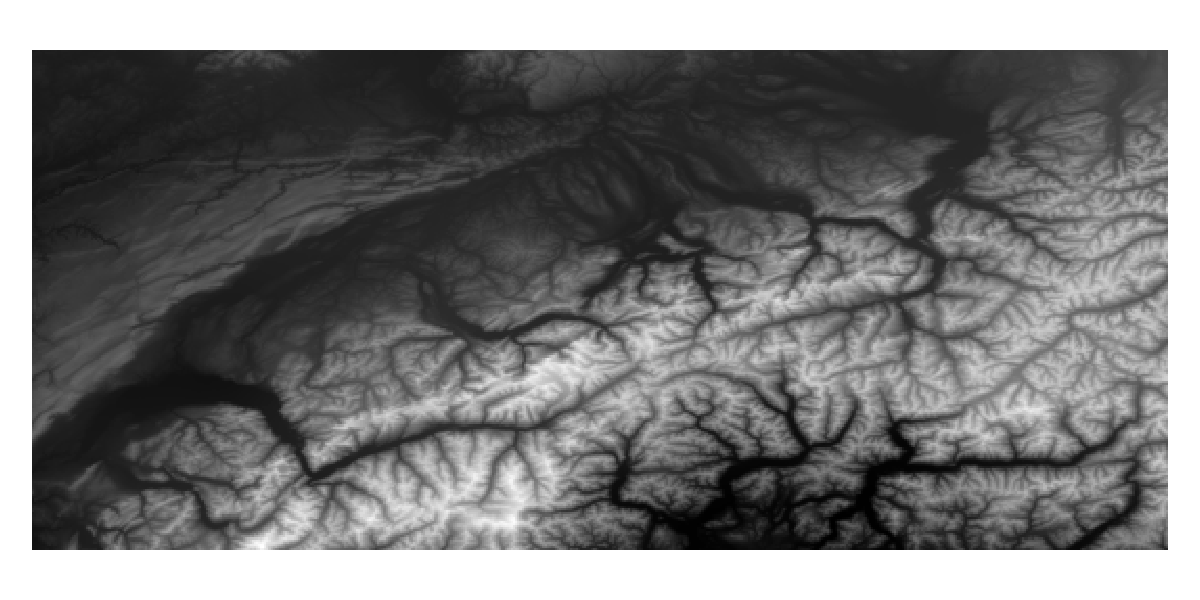
Code for the figure
f = Figure(; size = (600, 300))
ax = Axis(f[1, 1]; aspect = DataAspect())
heatmap!(ax, L[1]; colormap = :Greys)
hidespines!(ax)
hidedecorations!(ax)We can mask the entire vector of layers with the polygon we use to define our area of interest. This operation will modify the data in place, so we do not create additional objects.
mask!(L, aoi);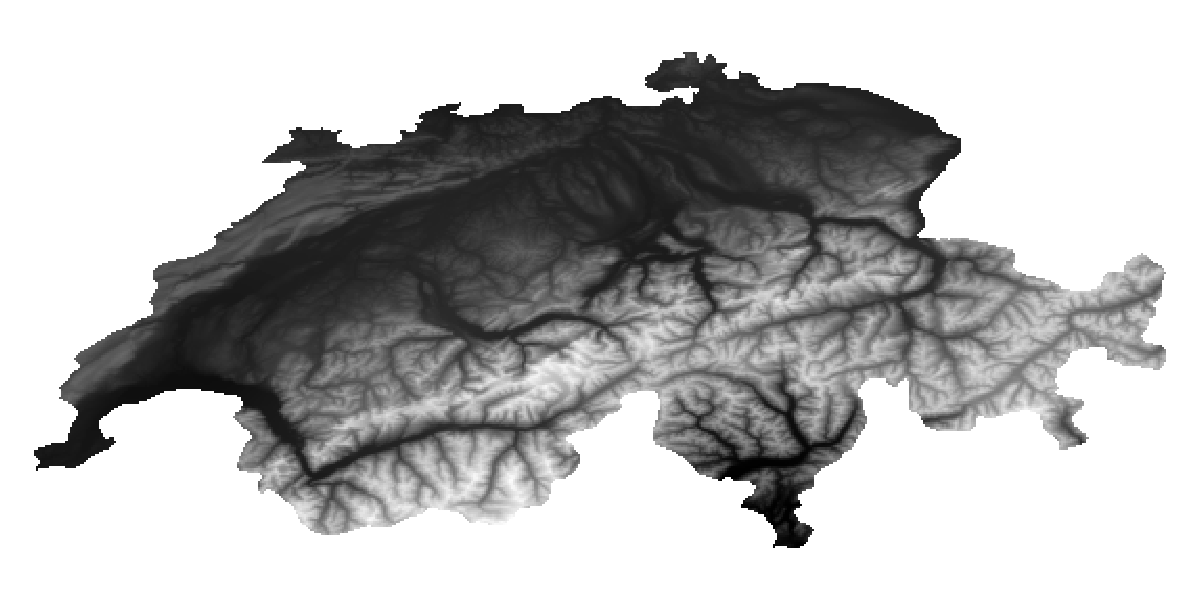
Code for the figure
f = Figure(; size = (600, 300))
ax = Axis(f[1, 1]; aspect = DataAspect())
heatmap!(ax, L[1]; colormap = :Greys)
hidespines!(ax)
hidedecorations!(ax)There is also a trim function, which will return a copy of the layers that are re-scaled so that they do not have empty rows/columns on either end. This is not required here, but may come in handy when doing more complex masking operations.
Masking layers
Many objects can be masked with polygons, and there are additional methods to mask layers as well. Both are documented in their vignettes. Masking a vector of layers is always faster than masking several layers in a row, because the mask is only calculated once and then re-used.
We will now grab a dataset from GBIF, which is a list of all observations of Turdus torquatus in Switzerland according to eBird. We will also grab a second dataset to test the model, which is the same species and country but uses iNaturalist data. Finally, we will get a representation of the
records = GBIF.download("10.15468/dl.wye52h")
test_records = GBIF.download("10.15468/dl.m6b3wg")
ouzel = taxon(first(entity(records)))GBIF taxon -- Turdus torquatusGBIF data
There are additional vignettes about retrieving data from the GBIF API, from GBIF downloads, or from local GBIF archives.
Getting the species as its own variable will allow us, among other things, to retrieve a silhouette from the Phylopic database. Note that it may not be the exact species, but Phylopic curates a taxonomy to return the closest matching organism. If your species is not in the Phylopic database, consider drawing the silhouette and contributing!
silhouette = Phylopic.imagesof(ouzel; items = 1)
Phylopic.attribution(silhouette)Image of Turdus migratorius provided by Andy Wilson
We can see the result of our data collection work thus far, by plotting the training and testing records on the region of interest, and adding a silhouette of the species alongside this map.
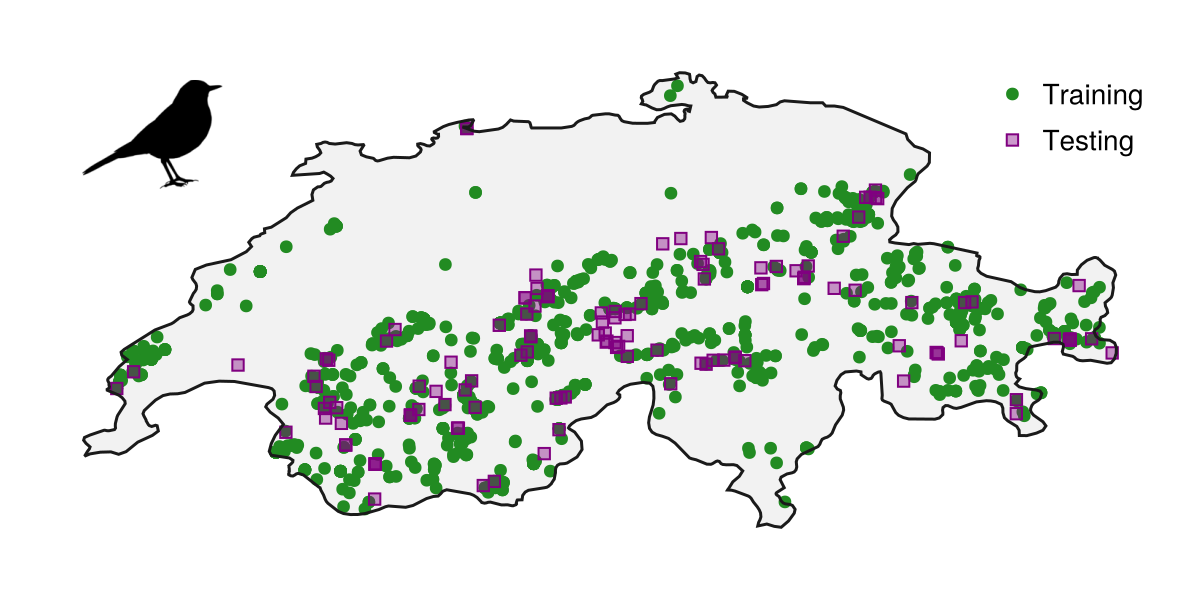
Code for the figure
f = Figure(; size = (600, 300))
ax = Axis(f[1, 1]; aspect = DataAspect())
poly!(ax, aoi; color = :grey95)
scatter!(ax, records; color = :forestgreen, label = "Training")
scatter!(
ax,
test_records;
color = (:purple, 0.4),
strokecolor = :purple,
strokewidth = 1,
marker = :rect,
markersize = 9,
label = "Testing",
)
hidedecorations!(ax)
hidespines!(ax)
lines!(ax, aoi; color = :grey10)
axislegend(ax; position = :rt, framevisible = false)
silhouetteplot!(
ax,
extent.left + 0.3,
extent.top - 0.2,
silhouette;
markersize = 70,
color = :black,
)At this point, we have collected all of the actual data we can, and it is time to generate pseudo-absences in order to have all of the material to train our model.
Pseudo-absence generation
We start by preparing a layer with presence data. This operation will ensure that all of the observations that share a raster cell will only be counted once. Note that for now, we will only work on the training records (from eBird).
presencelayer = mask(first(L), Occurrences(mask(records, aoi)))🗺️ A 239 × 543 layer with 70285 Bool cells
Spatial Reference System: +proj=longlat +datum=WGS84 +no_defsThe next step is to generate a pseudo-absence mask. We will sample according to the square root of the distance away from a known observation, but never sample a pseudo-absence less than 4km away from an observation, and never further away than 30km..
background = sqrt.(pseudoabsencemask(DistanceToEvent, presencelayer))🗺️ A 239 × 543 layer with 70285 Float64 cells
Spatial Reference System: +proj=longlat +datum=WGS84 +no_defsWithin this layer of possible location of absences, we will sample at random twice as many pseudo-absences cells as we have presence cells in the presence layer:
bgpoints = backgroundpoints(nodata(background, x -> !(sqrt(4.) <= x <= sqrt(30.))), 2sum(presencelayer))🗺️ A 239 × 543 layer with 44754 Bool cells
Spatial Reference System: +proj=longlat +datum=WGS84 +no_defsMore on pseudo-absences
There is a vignette with all the pseudo-absence generation methods. A lot of them are built into the package, and adding custom ones is not very difficult!
We can take a minute to visualize the dataset, as well as the location of presences and pseudo-absences:
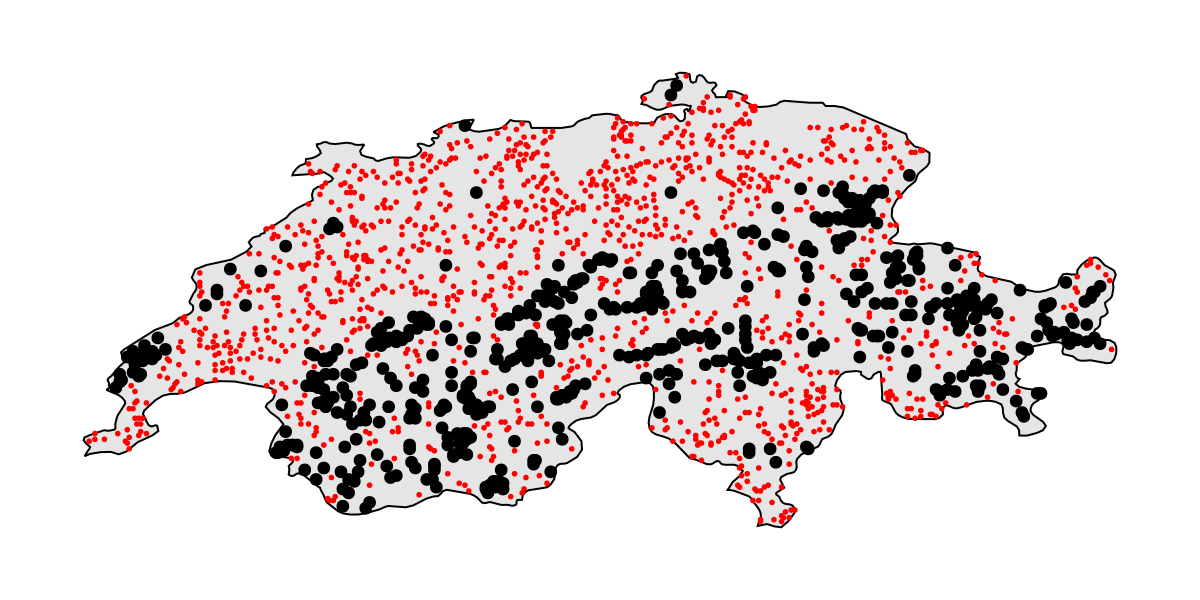
Code for the figure
f = Figure(; size = (600, 300))
ax = Axis(f[1, 1]; aspect = DataAspect())
poly!(ax, aoi; color = :grey90, strokecolor = :black, strokewidth = 1)
scatter!(ax, presencelayer; color = :black)
scatter!(ax, bgpoints; color = :red, markersize = 4)
hidedecorations!(ax)
hidespines!(ax)We now have enough information to set-up the actual species distribution model!
Setting up the model
We will use a model with a PCA for variable transformation, and then use logistic regression with all interactions between variables (i.e. both xᵢ² and xᵢxⱼ) to make the actual prediction. Note that we build this model by passing first the data transformation and the classifier, and then the vector of covariates, the layer with the presences, and the layer with the absences.
model = SDM(PCATransform, Logistic, L, presencelayer, bgpoints)❎ PCATransform → Logistic → P(x) ≥ 0.5 🗺️More on models
There is a vignette on model training that covers all of the options, as well as the method to generate ensemble and boosted models. It is is strongly advised to read it after this one. Or do you prefer to use Maxent? You can!.
We can tweak the hyperparameters of this model. We can check the default values for this classifier:
hyperparameters(classifier(model))Dict{Symbol, Any} with 6 entries:
:verbose => false
:interactions => :all
:λ => 0.1
:verbosity => 100
:epochs => 2000
:η => 0.01Hyper-parameters
There is a vignette on hyper-parameters that covers all of the functions to get and modify the hyper-parameters of a model.
We will set different hyper-parameters, to ensure that the learning rate is slower, the regularization is higher, and we do more training epochs:
hyperparameters!(classifier(model), :η, 1e-4);
hyperparameters!(classifier(model), :λ, 2e-1);
hyperparameters!(classifier(model), :epochs, 10_000);Note that, because we are generating the SDM from a series of layers, it is georeferenced. A georeferenced model can be plotted directly, as well as used to define spatial cross-validation blocks. We can confirm that the model is georeferenced:
isgeoreferenced(model)trueOur model is now ready to be trained. Before we train it, we will perform cross-validation and variable selection.
Variable selection
In this section, we will use cross-validation to identify the optimal set of variables for the model. The optimal set of variables is defined as the one that gives the best performance on average over the validation sets.
More on cross-validation
Both cross-validation and feature selection are covered in their own vignettes.
We start by splitting the dataset into training and validation data. We will use k-fold cross-validation using five splits (the default is ten). By default, the splits are always balanced, so that the training and validation data have the same prevalence of presences.
folds = kfold(model; k=5)5-element Vector{Tuple{Vector{Int64}, Vector{Int64}}}:
([1, 2, 3, 4, 6, 7, 9, 10, 11, 12, 14, 15, 16, 17, 18, 19, 20, 21, 23, 24, 25, 27, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 47, 48, 49, 50, 52, 55, 57, 58, 59, 60, 62, 65, 66, 67, 69, 70, 71, 72, 73, 74, 75, 76, 78, 79, 81, 82, 83, 84, 85, 86, 87, 88, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, 100, 102, 103, 104, 105, 106, 107, 108, 109, 110, 111, 112, 113, 114, 116, 117, 118, 119, 120, 121, 122, 124, 125, 126, 127, 128, 129, 130, 131, 132, 134, 136, 137, 138, 139, 140, 141, 142, 143, 144, 145, 146, 147, 148, 149, 151, 152, 153, 154, 155, 156, 159, 160, 161, 163, 164, 165, 166, 167, 168, 170, 175, 176, 178, 180, 182, 183, 184, 185, 186, 189, 190, 191, 192, 193, 194, 195, 196, 199, 200, 201, 202, 205, 206, 207, 208, 209, 210, 212, 213, 214, 215, 216, 218, 219, 220, 221, 223, 224, 225, 226, 227, 229, 230, 231, 232, 234, 236, 237, 239, 240, 241, 243, 244, 245, 246, 247, 248, 249, 250, 251, 252, 253, 255, 256, 257, 259, 261, 262, 263, 264, 265, 266, 267, 269, 270, 271, 272, 274, 275, 276, 278, 279, 280, 281, 282, 283, 284, 287, 288, 289, 290, 291, 292, 293, 294, 295, 296, 297, 298, 299, 300, 301, 302, 303, 304, 306, 307, 308, 310, 311, 312, 314, 316, 317, 318, 319, 320, 322, 323, 325, 326, 327, 328, 329, 330, 331, 332, 333, 334, 335, 336, 337, 341, 342, 343, 345, 346, 347, 348, 349, 350, 351, 353, 355, 356, 357, 358, 359, 360, 362, 363, 365, 367, 368, 370, 371, 372, 373, 374, 375, 376, 378, 380, 382, 384, 386, 388, 389, 390, 391, 392, 393, 394, 396, 399, 401, 402, 404, 405, 406, 407, 408, 409, 410, 411, 412, 413, 414, 415, 416, 417, 420, 421, 422, 423, 424, 425, 426, 427, 428, 429, 430, 431, 433, 434, 435, 436, 437, 438, 439, 440, 441, 442, 443, 444, 445, 446, 447, 449, 450, 451, 452, 453, 454, 456, 457, 458, 459, 460, 461, 462, 463, 464, 466, 467, 468, 469, 470, 473, 474, 476, 477, 478, 479, 480, 481, 482, 484, 487, 488, 489, 490, 492, 493, 495, 496, 497, 499, 501, 504, 506, 507, 508, 511, 513, 514, 515, 517, 518, 519, 520, 521, 522, 525, 526, 527, 528, 529, 530, 531, 532, 534, 535, 536, 537, 538, 539, 540, 541, 543, 544, 546, 547, 548, 549, 550, 551, 552, 553, 554, 555, 556, 557, 558, 559, 560, 561, 562, 563, 564, 565, 566, 567, 568, 569, 570, 571, 573, 574, 576, 577, 578, 579, 580, 581, 582, 583, 584, 585, 587, 588, 589, 591, 592, 594, 595, 596, 597, 599, 600, 601, 602, 603, 604, 605, 608, 610, 611, 612, 613, 615, 616, 617, 619, 620, 621, 622, 623, 624, 625, 626, 628, 629, 630, 631, 633, 634, 635, 637, 638, 639, 641, 642, 644, 646, 647, 648, 649, 650, 651, 652, 653, 654, 656, 657, 659, 660, 661, 662, 664, 665, 666, 668, 669, 670, 672, 673, 674, 676, 678, 679, 680, 681, 682, 683, 684, 685, 688, 689, 690, 692, 693, 694, 695, 696, 697, 698, 699, 701, 702, 703, 704, 706, 708, 709, 711, 712, 713, 714, 715, 717, 718, 719, 720, 722, 725, 726, 727, 728, 729, 730, 731, 732, 733, 734, 735, 736, 737, 738, 740, 742, 744, 745, 746, 747, 748, 750, 751, 752, 753, 754, 755, 756, 759, 760, 761, 762, 764, 765, 766, 767, 768, 770, 772, 774, 775, 776, 777, 778, 779, 780, 782, 783, 784, 785, 787, 788, 789, 790, 791, 792, 793, 794, 795, 796, 797, 798, 799, 800, 801, 802, 803, 804, 805, 806, 807, 808, 809, 810, 811, 812, 813, 816, 820, 821, 822, 823, 824, 825, 826, 827, 828, 829, 830, 831, 832, 834, 835, 836, 837, 838, 839, 840, 841, 842, 843, 844, 845, 846, 847, 848, 849, 850, 851, 853, 854, 855, 856, 858, 859, 861, 863, 864, 865, 866, 867, 868, 869, 870, 871, 872, 873, 874, 875, 878, 879, 881, 882, 883, 884, 886, 887, 888, 889, 891, 892, 893, 894, 895, 896, 897, 898, 899, 900, 901, 902, 904, 907, 908, 909, 910, 912, 913, 915, 916, 918, 919, 921, 922, 923, 924, 926, 927, 928, 929, 930, 931, 932, 933, 934, 936, 937, 938, 939, 940, 941, 942, 943, 944, 945, 946, 948, 949, 950, 952, 953, 954, 955, 956, 957, 958, 961, 962, 963, 964, 965, 966, 967, 968, 969, 970, 971, 972, 973, 974, 975, 976, 977, 978, 979, 980, 981, 982, 983, 984, 986, 987, 988, 990, 993, 994, 996, 997, 1000, 1002, 1003, 1004, 1005, 1007, 1008, 1009, 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1017, 1019, 1020, 1021, 1022, 1023, 1025, 1026, 1027, 1028, 1029, 1030, 1031, 1033, 1034, 1035, 1036, 1037, 1038, 1039, 1041, 1042, 1043, 1045, 1046, 1048, 1049, 1050, 1051, 1053, 1054, 1055, 1056, 1057, 1058, 1061, 1062, 1063, 1064, 1066, 1067, 1068, 1069, 1070, 1071, 1072, 1073, 1074, 1075, 1076, 1077, 1078, 1081, 1082, 1083, 1084, 1085, 1086, 1087, 1089, 1090, 1091, 1093, 1094, 1095, 1096, 1097, 1098, 1099, 1100, 1101, 1102, 1103, 1104, 1106, 1107, 1108, 1109, 1110, 1112, 1113, 1114, 1116, 1117, 1119, 1120, 1121, 1122, 1123, 1124, 1125, 1126, 1127, 1128, 1129, 1130, 1131, 1132, 1133, 1134, 1136, 1137, 1138, 1139, 1140, 1141, 1142, 1143, 1144, 1145, 1146, 1147, 1149, 1150, 1151, 1152, 1154, 1156, 1157, 1159, 1160, 1161, 1162, 1164, 1167, 1169, 1170, 1171, 1172, 1174, 1175, 1177, 1178, 1179, 1180, 1181, 1184, 1185, 1186, 1187, 1188, 1189, 1190, 1191, 1192, 1193, 1194, 1196, 1197, 1198, 1199, 1202, 1205, 1206, 1207, 1209, 1210, 1211, 1212, 1213, 1214, 1215, 1217, 1218, 1219, 1220, 1221, 1222, 1223, 1225, 1226, 1227, 1228, 1229, 1230, 1231, 1232, 1233, 1234, 1235, 1236, 1237, 1238, 1239, 1240, 1241, 1242, 1243, 1245, 1246, 1248, 1249, 1250, 1251, 1252, 1253, 1254, 1255, 1256, 1257, 1259, 1260, 1263, 1265, 1266, 1267, 1269, 1270, 1271, 1272, 1273, 1274, 1275, 1276, 1277, 1278, 1279, 1280, 1281, 1282, 1284, 1285, 1288, 1289, 1291, 1292, 1293, 1294, 1295, 1296, 1298, 1299, 1301, 1302, 1303, 1305, 1306, 1307, 1308, 1309, 1310, 1311, 1313, 1314, 1315, 1318, 1320, 1321, 1323, 1324, 1326, 1327, 1328, 1329, 1330, 1332, 1333, 1334, 1335, 1336, 1337, 1338, 1339, 1341, 1342, 1343, 1344, 1345, 1346, 1347, 1348, 1349, 1350, 1351, 1352, 1353, 1354, 1355, 1356, 1357, 1359, 1360, 1361, 1362, 1363, 1364, 1365, 1366, 1367, 1368, 1369, 1370, 1371, 1373, 1375, 1376, 1377, 1378, 1379, 1380, 1381, 1382, 1384, 1385, 1386, 1387, 1388, 1389, 1390, 1391, 1393, 1394, 1395, 1396, 1397, 1398, 1399, 1401, 1402, 1405, 1408, 1410, 1412, 1413, 1414, 1415, 1416, 1419, 1420, 1422, 1424, 1425, 1426, 1427, 1428, 1429, 1431, 1432, 1433, 1434, 1435, 1436, 1437, 1438, 1439, 1440, 1441, 1442, 1444, 1445, 1447, 1448, 1449, 1450, 1452, 1454, 1455, 1457, 1458, 1459, 1460, 1461, 1462, 1463, 1464, 1465, 1466, 1467, 1468, 1469, 1470, 1471, 1474, 1475, 1476, 1477, 1478, 1479, 1480, 1481, 1482, 1484, 1485, 1486, 1487, 1488, 1489, 1490, 1491, 1492, 1494, 1495, 1496, 1498, 1499, 1501, 1502, 1504, 1505, 1506, 1507, 1510, 1511, 1512, 1513, 1514, 1515, 1516, 1517, 1518, 1520, 1522, 1523, 1525, 1526, 1527, 1528, 1529, 1531, 1532, 1534, 1535, 1536, 1537, 1538, 1539, 1540, 1541, 1543, 1544, 1545, 1547, 1548, 1549, 1550, 1551, 1553, 1554, 1555, 1556, 1557, 1558, 1559, 1560, 1561, 1562, 1563, 1564, 1565, 1566, 1569, 1570, 1572, 1573, 1574, 1575, 1576, 1578, 1579, 1580, 1581, 1582, 1583, 1584, 1587, 1588, 1589, 1590, 1591, 1593, 1594, 1596, 1597, 1598, 1600, 1601, 1603, 1604, 1606, 1607, 1608, 1609, 1610, 1611, 1612, 1613, 1614, 1615, 1616, 1617, 1618, 1619, 1621, 1622, 1623, 1624, 1625, 1627, 1628, 1629, 1630, 1631, 1632, 1636, 1639, 1640, 1641, 1642, 1643, 1646, 1647, 1648, 1649, 1653, 1655, 1656, 1657, 1658, 1660, 1661, 1662, 1663, 1665, 1666, 1667, 1668, 1669, 1671, 1672, 1673, 1674, 1676, 1677, 1678, 1679, 1680, 1681, 1682, 1683, 1684, 1685, 1686, 1688, 1689, 1690, 1692, 1693, 1694, 1695, 1696, 1697, 1699, 1700, 1701, 1702, 1704, 1705, 1707, 1708, 1709, 1711, 1712, 1713, 1714, 1715, 1716, 1717, 1719, 1720, 1721, 1723, 1724, 1725, 1726, 1727, 1728, 1729, 1731, 1733, 1734, 1736, 1737, 1738, 1740, 1741, 1742, 1745, 1746, 1747, 1748, 1749, 1750, 1752, 1753, 1754, 1755, 1756, 1757, 1758, 1760, 1762, 1763, 1764, 1765, 1766, 1767, 1768, 1769, 1770, 1771, 1772, 1774, 1775, 1776, 1777, 1778, 1779, 1780, 1781, 1782, 1783, 1784, 1785, 1786, 1787, 1788, 1790, 1791, 1792, 1793, 1795, 1797, 1799, 1803, 1805, 1806, 1808, 1809, 1810, 1811, 1815, 1816, 1817, 1819, 1820, 1821, 1822, 1823, 1824, 1826, 1830, 1832, 1833, 1834, 1835, 1836, 1837, 1838, 1839, 1840, 1841, 1843, 1845, 1846, 1848, 1849, 1851, 1852, 1855, 1856, 1857, 1858, 1859, 1860, 1861, 1863, 1864, 1866, 1867, 1868, 1869, 1870, 1871, 1872, 1873, 1876, 1877, 1878, 1879, 1880, 1881, 1882, 1887, 1888, 1889, 1890, 1891, 1896, 1898, 1900, 1901, 1902, 1903, 1904, 1905, 1908, 1910, 1912, 1913, 1914, 1915, 1916, 1917, 1918, 1921, 1922, 1924, 1925, 1926, 1927, 1928, 1929, 1930, 1931, 1932, 1934, 1935, 1936, 1937, 1939, 1940, 1941, 1943, 1944, 1945, 1946, 1948, 1949, 1950, 1951, 1952, 1953, 1954, 1956, 1958, 1959, 1960, 1961, 1962], [204, 516, 339, 379, 56, 45, 309, 383, 222, 115, 593, 658, 197, 315, 590, 77, 135, 465, 494, 188, 533, 64, 663, 471, 217, 395, 8, 509, 173, 233, 285, 61, 607, 181, 483, 403, 598, 133, 46, 5, 340, 524, 174, 618, 643, 313, 381, 606, 655, 80, 235, 366, 305, 387, 171, 385, 150, 364, 545, 361, 352, 228, 575, 172, 286, 268, 179, 51, 432, 609, 448, 472, 277, 512, 627, 68, 455, 101, 123, 485, 500, 324, 418, 397, 398, 369, 203, 645, 211, 273, 344, 89, 254, 158, 13, 26, 505, 258, 28, 419, 354, 503, 498, 632, 614, 523, 542, 510, 63, 54, 636, 475, 338, 572, 491, 377, 640, 162, 502, 22, 157, 238, 260, 486, 177, 400, 169, 321, 242, 586, 198, 53, 187, 1392, 1865, 1533, 1200, 723, 1183, 1261, 769, 1818, 1691, 1751, 1812, 1258, 1710, 758, 1814, 1509, 1418, 1224, 857, 947, 1040, 1807, 1885, 1626, 1586, 1650, 1216, 1652, 1176, 700, 1032, 903, 1503, 1018, 880, 1893, 1407, 1052, 951, 1358, 1135, 1894, 1406, 1262, 1732, 1047, 1148, 675, 1111, 1911, 1568, 1802, 1290, 959, 786, 1001, 818, 1268, 1247, 1909, 677, 1524, 890, 1453, 1287, 852, 1761, 1163, 885, 1403, 917, 1718, 1567, 1831, 876, 1168, 1472, 1735, 992, 1500, 1759, 1080, 1850, 1118, 1203, 1166, 1892, 1497, 1340, 1331, 1796, 1092, 1079, 1907, 1635, 1244, 1804, 1703, 999, 925, 1886, 716, 1322, 998, 1059, 1508, 1201, 1115, 1530, 721, 1842, 1519, 757, 749, 671, 1862, 771, 1182, 1483, 1827, 1325, 1947, 1546, 1844, 1744, 1906, 1633, 1165, 1493, 1599, 1317, 989, 1158, 833, 763, 991, 1938, 1374, 1899, 877, 1637, 1638, 1595, 1155, 1789, 1024, 815, 1698, 1446, 1813, 1739, 1634, 1542, 1722, 1895, 1875, 1874, 1651, 1659, 1060, 906, 1645, 1319, 1264, 1195, 1801, 773, 1173, 741, 1602, 935, 1853, 819, 1825, 1571, 1919, 1153, 1297, 1923, 1088, 1105, 1065, 1430, 1423, 1521, 1883, 781, 1828, 1884, 960, 1421, 1854, 1706, 1300, 914, 1920, 1743, 814, 1312, 1847, 1829, 707, 1404, 1897, 1204, 1585, 817, 1644, 1044, 691, 1687, 995, 1283, 1304, 1443, 1800, 1316, 743, 1794, 1664, 1605, 1620, 1730, 1577, 860, 687, 905, 911, 862, 1456, 724, 1473, 710, 1592, 985, 1654, 739, 1400, 1208, 1670, 1552, 686, 1675, 920, 1006, 1417, 1409, 1286, 1933, 1955, 1372, 705, 1798, 1451, 1773, 1411, 667, 1942, 1957, 1383])
([1, 2, 3, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 36, 37, 38, 39, 40, 41, 44, 45, 46, 47, 48, 49, 50, 51, 53, 54, 56, 58, 60, 61, 63, 64, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 85, 87, 89, 90, 92, 93, 95, 97, 98, 99, 100, 101, 102, 103, 105, 106, 107, 109, 110, 111, 112, 113, 114, 115, 116, 117, 120, 121, 122, 123, 125, 126, 127, 129, 130, 131, 132, 133, 135, 136, 137, 138, 139, 140, 141, 144, 146, 147, 148, 149, 150, 151, 152, 153, 154, 155, 156, 157, 158, 159, 160, 162, 164, 165, 166, 167, 168, 169, 170, 171, 172, 173, 174, 176, 177, 178, 179, 180, 181, 182, 183, 184, 186, 187, 188, 189, 190, 191, 192, 193, 194, 195, 196, 197, 198, 200, 201, 203, 204, 205, 207, 209, 210, 211, 212, 213, 214, 216, 217, 218, 220, 222, 223, 224, 227, 228, 229, 231, 232, 233, 234, 235, 236, 237, 238, 240, 241, 242, 244, 245, 246, 247, 248, 249, 250, 251, 252, 253, 254, 256, 257, 258, 259, 260, 261, 262, 264, 266, 267, 268, 269, 270, 271, 272, 273, 274, 275, 276, 277, 278, 279, 280, 281, 282, 283, 284, 285, 286, 287, 288, 289, 291, 292, 293, 295, 296, 297, 298, 299, 300, 301, 302, 303, 304, 305, 307, 308, 309, 310, 311, 312, 313, 314, 315, 317, 318, 319, 320, 321, 322, 324, 325, 326, 327, 328, 330, 331, 332, 333, 335, 336, 337, 338, 339, 340, 341, 342, 344, 345, 346, 348, 349, 351, 352, 353, 354, 355, 357, 358, 359, 360, 361, 363, 364, 365, 366, 367, 369, 370, 371, 372, 375, 376, 377, 378, 379, 380, 381, 382, 383, 384, 385, 386, 387, 388, 390, 392, 393, 394, 395, 396, 397, 398, 400, 403, 404, 405, 406, 409, 411, 412, 413, 414, 416, 417, 418, 419, 421, 422, 426, 428, 431, 432, 433, 434, 435, 438, 440, 441, 442, 443, 445, 446, 448, 449, 450, 451, 452, 453, 454, 455, 456, 457, 460, 463, 464, 465, 466, 467, 468, 469, 471, 472, 473, 474, 475, 476, 477, 478, 479, 481, 482, 483, 484, 485, 486, 487, 488, 489, 491, 492, 494, 495, 496, 497, 498, 499, 500, 501, 502, 503, 504, 505, 506, 509, 510, 511, 512, 513, 514, 515, 516, 517, 519, 520, 521, 523, 524, 525, 526, 527, 528, 529, 530, 531, 532, 533, 535, 536, 537, 538, 539, 542, 545, 546, 547, 549, 550, 551, 552, 554, 555, 556, 557, 558, 559, 560, 561, 562, 563, 565, 567, 569, 571, 572, 573, 574, 575, 577, 578, 579, 580, 581, 585, 586, 587, 589, 590, 591, 592, 593, 595, 597, 598, 599, 601, 604, 605, 606, 607, 608, 609, 613, 614, 615, 618, 619, 621, 622, 623, 626, 627, 628, 629, 630, 631, 632, 633, 634, 635, 636, 638, 639, 640, 641, 643, 645, 646, 647, 648, 649, 650, 651, 655, 657, 658, 659, 660, 663, 664, 665, 667, 668, 669, 670, 671, 673, 675, 677, 678, 680, 681, 682, 684, 686, 687, 688, 691, 692, 693, 695, 696, 700, 703, 704, 705, 706, 707, 708, 710, 711, 712, 713, 714, 715, 716, 717, 718, 719, 720, 721, 723, 724, 725, 726, 728, 729, 730, 731, 732, 733, 734, 735, 736, 737, 739, 741, 743, 744, 745, 746, 747, 748, 749, 751, 752, 753, 754, 755, 756, 757, 758, 759, 760, 761, 762, 763, 764, 765, 766, 768, 769, 771, 773, 774, 776, 778, 779, 780, 781, 782, 784, 785, 786, 787, 788, 789, 790, 793, 794, 795, 797, 798, 799, 800, 803, 804, 806, 807, 808, 809, 811, 813, 814, 815, 816, 817, 818, 819, 820, 821, 824, 825, 826, 827, 829, 830, 832, 833, 834, 835, 836, 838, 839, 840, 842, 843, 844, 846, 847, 848, 849, 850, 851, 852, 853, 855, 857, 859, 860, 861, 862, 863, 864, 865, 866, 867, 868, 869, 870, 874, 875, 876, 877, 878, 880, 881, 882, 883, 884, 885, 886, 887, 889, 890, 891, 892, 894, 895, 896, 898, 900, 902, 903, 904, 905, 906, 908, 910, 911, 912, 913, 914, 915, 916, 917, 919, 920, 921, 922, 923, 924, 925, 926, 927, 928, 929, 931, 932, 933, 935, 936, 937, 938, 940, 941, 942, 943, 944, 945, 946, 947, 948, 949, 951, 953, 954, 955, 956, 957, 959, 960, 961, 962, 963, 966, 967, 968, 969, 972, 973, 975, 976, 977, 978, 979, 982, 983, 984, 985, 986, 989, 990, 991, 992, 993, 994, 995, 996, 997, 998, 999, 1000, 1001, 1002, 1004, 1006, 1008, 1011, 1012, 1013, 1014, 1015, 1016, 1017, 1018, 1019, 1020, 1021, 1022, 1023, 1024, 1025, 1027, 1028, 1029, 1030, 1031, 1032, 1033, 1035, 1036, 1037, 1038, 1039, 1040, 1042, 1044, 1046, 1047, 1048, 1049, 1051, 1052, 1053, 1054, 1055, 1056, 1059, 1060, 1061, 1063, 1064, 1065, 1066, 1067, 1069, 1070, 1071, 1072, 1073, 1075, 1077, 1078, 1079, 1080, 1082, 1083, 1084, 1085, 1086, 1087, 1088, 1089, 1090, 1091, 1092, 1094, 1095, 1097, 1098, 1100, 1101, 1104, 1105, 1106, 1107, 1108, 1110, 1111, 1112, 1113, 1114, 1115, 1118, 1119, 1120, 1121, 1122, 1123, 1124, 1125, 1126, 1127, 1128, 1130, 1131, 1132, 1133, 1134, 1135, 1136, 1138, 1139, 1140, 1141, 1142, 1143, 1144, 1145, 1146, 1147, 1148, 1149, 1150, 1151, 1152, 1153, 1154, 1155, 1156, 1158, 1159, 1160, 1161, 1162, 1163, 1164, 1165, 1166, 1167, 1168, 1169, 1171, 1172, 1173, 1175, 1176, 1178, 1179, 1180, 1182, 1183, 1185, 1186, 1187, 1188, 1189, 1190, 1192, 1193, 1195, 1196, 1197, 1199, 1200, 1201, 1202, 1203, 1204, 1205, 1206, 1207, 1208, 1210, 1211, 1212, 1213, 1214, 1215, 1216, 1217, 1218, 1219, 1220, 1221, 1222, 1223, 1224, 1225, 1226, 1227, 1228, 1229, 1230, 1231, 1232, 1233, 1235, 1238, 1239, 1240, 1242, 1244, 1245, 1246, 1247, 1248, 1249, 1250, 1254, 1256, 1257, 1258, 1259, 1261, 1262, 1264, 1265, 1266, 1267, 1268, 1269, 1270, 1271, 1272, 1273, 1274, 1275, 1276, 1277, 1278, 1279, 1281, 1283, 1284, 1286, 1287, 1289, 1290, 1292, 1293, 1294, 1295, 1296, 1297, 1300, 1301, 1302, 1303, 1304, 1305, 1306, 1307, 1308, 1309, 1310, 1311, 1312, 1315, 1316, 1317, 1318, 1319, 1320, 1322, 1323, 1324, 1325, 1326, 1328, 1329, 1330, 1331, 1332, 1333, 1334, 1335, 1336, 1338, 1340, 1341, 1344, 1345, 1346, 1347, 1349, 1351, 1353, 1355, 1357, 1358, 1359, 1361, 1362, 1364, 1366, 1367, 1368, 1369, 1370, 1371, 1372, 1373, 1374, 1375, 1376, 1377, 1378, 1379, 1380, 1381, 1382, 1383, 1384, 1386, 1387, 1388, 1390, 1391, 1392, 1393, 1395, 1396, 1397, 1398, 1400, 1401, 1402, 1403, 1404, 1405, 1406, 1407, 1409, 1410, 1411, 1412, 1413, 1414, 1416, 1417, 1418, 1420, 1421, 1422, 1423, 1424, 1425, 1426, 1430, 1431, 1432, 1433, 1434, 1435, 1436, 1437, 1438, 1439, 1441, 1442, 1443, 1444, 1445, 1446, 1448, 1449, 1450, 1451, 1452, 1453, 1454, 1455, 1456, 1458, 1459, 1461, 1463, 1464, 1468, 1469, 1471, 1472, 1473, 1474, 1476, 1478, 1479, 1480, 1483, 1484, 1485, 1486, 1488, 1489, 1492, 1493, 1494, 1496, 1497, 1498, 1499, 1500, 1503, 1504, 1505, 1506, 1507, 1508, 1509, 1510, 1511, 1512, 1513, 1514, 1516, 1517, 1519, 1520, 1521, 1522, 1523, 1524, 1525, 1526, 1527, 1528, 1529, 1530, 1533, 1534, 1535, 1536, 1537, 1538, 1539, 1541, 1542, 1543, 1544, 1545, 1546, 1547, 1548, 1549, 1551, 1552, 1553, 1558, 1559, 1560, 1561, 1563, 1565, 1566, 1567, 1568, 1569, 1570, 1571, 1572, 1573, 1574, 1575, 1576, 1577, 1579, 1580, 1581, 1582, 1583, 1584, 1585, 1586, 1587, 1588, 1589, 1590, 1591, 1592, 1593, 1595, 1596, 1597, 1598, 1599, 1600, 1601, 1602, 1603, 1604, 1605, 1606, 1607, 1608, 1609, 1610, 1611, 1612, 1614, 1615, 1616, 1617, 1618, 1619, 1620, 1623, 1624, 1625, 1626, 1628, 1629, 1630, 1632, 1633, 1634, 1635, 1636, 1637, 1638, 1639, 1640, 1642, 1643, 1644, 1645, 1646, 1648, 1649, 1650, 1651, 1652, 1653, 1654, 1655, 1657, 1658, 1659, 1660, 1661, 1663, 1664, 1666, 1667, 1669, 1670, 1673, 1674, 1675, 1676, 1679, 1680, 1683, 1684, 1686, 1687, 1689, 1691, 1693, 1695, 1696, 1697, 1698, 1699, 1700, 1701, 1702, 1703, 1704, 1705, 1706, 1708, 1710, 1713, 1714, 1715, 1716, 1717, 1718, 1719, 1720, 1721, 1722, 1724, 1726, 1727, 1728, 1729, 1730, 1731, 1732, 1733, 1735, 1736, 1737, 1738, 1739, 1740, 1741, 1742, 1743, 1744, 1745, 1746, 1748, 1749, 1750, 1751, 1752, 1753, 1756, 1758, 1759, 1760, 1761, 1762, 1764, 1765, 1766, 1767, 1768, 1769, 1770, 1771, 1772, 1773, 1774, 1775, 1776, 1777, 1778, 1779, 1780, 1781, 1782, 1783, 1784, 1785, 1786, 1789, 1790, 1791, 1793, 1794, 1796, 1798, 1800, 1801, 1802, 1804, 1805, 1807, 1808, 1809, 1810, 1812, 1813, 1814, 1815, 1816, 1817, 1818, 1819, 1820, 1824, 1825, 1826, 1827, 1828, 1829, 1830, 1831, 1833, 1834, 1838, 1839, 1840, 1841, 1842, 1844, 1845, 1846, 1847, 1849, 1850, 1851, 1852, 1853, 1854, 1856, 1858, 1862, 1863, 1864, 1865, 1866, 1867, 1868, 1869, 1870, 1871, 1872, 1873, 1874, 1875, 1876, 1877, 1878, 1881, 1882, 1883, 1884, 1885, 1886, 1888, 1889, 1890, 1891, 1892, 1893, 1894, 1895, 1897, 1899, 1901, 1902, 1904, 1906, 1907, 1908, 1909, 1910, 1911, 1912, 1915, 1916, 1917, 1918, 1919, 1920, 1921, 1922, 1923, 1924, 1927, 1928, 1929, 1930, 1931, 1932, 1933, 1934, 1935, 1936, 1937, 1938, 1939, 1940, 1942, 1943, 1944, 1946, 1947, 1948, 1949, 1950, 1951, 1952, 1953, 1954, 1955, 1956, 1957, 1958, 1959, 1960, 1961, 1962], [343, 215, 108, 617, 86, 91, 461, 43, 59, 57, 145, 611, 399, 459, 368, 199, 423, 583, 88, 347, 653, 588, 323, 65, 429, 652, 206, 480, 644, 427, 356, 564, 219, 424, 263, 35, 94, 600, 334, 493, 226, 239, 221, 490, 540, 656, 661, 620, 294, 161, 401, 4, 544, 408, 402, 518, 522, 602, 654, 553, 124, 543, 415, 624, 96, 642, 458, 470, 534, 568, 637, 185, 84, 142, 625, 134, 306, 437, 42, 118, 202, 436, 163, 444, 329, 439, 541, 508, 373, 255, 616, 430, 391, 52, 62, 208, 316, 290, 612, 570, 596, 566, 243, 603, 128, 548, 425, 230, 350, 55, 362, 16, 584, 15, 143, 420, 576, 447, 104, 374, 265, 610, 462, 594, 119, 662, 507, 175, 410, 582, 407, 389, 225, 1010, 1057, 1313, 1823, 1836, 1879, 1117, 1282, 1285, 666, 1723, 685, 1243, 901, 970, 871, 1556, 1900, 742, 858, 698, 770, 1003, 980, 1356, 772, 1236, 1415, 1184, 802, 1194, 738, 1668, 1198, 1465, 1855, 1822, 1678, 699, 1389, 1251, 822, 1631, 981, 907, 1365, 1482, 1685, 1725, 1763, 676, 987, 1487, 1076, 1857, 1460, 1898, 1613, 974, 1671, 879, 1050, 1348, 1757, 1447, 1540, 1835, 1709, 1477, 854, 1707, 888, 1945, 1837, 1495, 1321, 1288, 1093, 1481, 1363, 1555, 1253, 1457, 777, 1427, 1299, 1803, 1594, 1501, 1502, 1564, 1440, 1116, 1041, 930, 1252, 1941, 1314, 1081, 1099, 1665, 958, 805, 1688, 1788, 1103, 1692, 1467, 1880, 939, 683, 1747, 918, 792, 1007, 1177, 1677, 1129, 873, 1034, 1181, 1298, 1799, 1795, 1712, 1342, 1280, 1518, 831, 690, 727, 775, 689, 1672, 988, 1241, 1806, 1811, 1622, 1343, 1058, 697, 965, 1337, 1096, 812, 801, 1009, 1662, 1260, 1005, 845, 1354, 1068, 1926, 1859, 701, 1550, 1754, 1903, 1711, 674, 1621, 1234, 1755, 810, 1327, 1263, 952, 1913, 1026, 1137, 1157, 1792, 1408, 1429, 1787, 1462, 1352, 1821, 1860, 1360, 1627, 1515, 1394, 1291, 1466, 672, 828, 856, 740, 1209, 1578, 893, 694, 1562, 1861, 964, 1045, 1887, 872, 1532, 971, 837, 1797, 1074, 1102, 1109, 1255, 783, 1470, 1170, 709, 679, 1531, 1656, 1490, 1848, 1385, 1681, 823, 1641, 1399, 1191, 1914, 791, 1694, 1905, 899, 767, 909, 1843, 1690, 1237, 1174, 1557, 1062, 934, 796, 1491, 722, 1734, 897, 950, 1682, 1647, 1419, 1428, 1554, 1350, 1475, 702, 1832, 1896, 841, 1339, 1925, 1043, 750])
([3, 4, 5, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 18, 19, 20, 22, 23, 24, 25, 26, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 40, 42, 43, 45, 46, 47, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 73, 77, 78, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 96, 97, 98, 99, 100, 101, 102, 103, 104, 105, 106, 107, 108, 111, 112, 113, 114, 115, 116, 118, 119, 120, 122, 123, 124, 125, 127, 128, 129, 130, 131, 133, 134, 135, 138, 139, 140, 141, 142, 143, 145, 146, 147, 148, 149, 150, 151, 153, 155, 156, 157, 158, 159, 160, 161, 162, 163, 164, 165, 166, 168, 169, 170, 171, 172, 173, 174, 175, 177, 179, 180, 181, 182, 183, 185, 187, 188, 189, 190, 191, 192, 193, 196, 197, 198, 199, 200, 201, 202, 203, 204, 205, 206, 208, 211, 212, 213, 215, 216, 217, 218, 219, 220, 221, 222, 223, 224, 225, 226, 227, 228, 230, 231, 232, 233, 234, 235, 236, 237, 238, 239, 241, 242, 243, 244, 246, 249, 250, 251, 253, 254, 255, 258, 259, 260, 261, 262, 263, 264, 265, 266, 267, 268, 270, 271, 272, 273, 274, 275, 276, 277, 278, 279, 280, 282, 284, 285, 286, 289, 290, 291, 292, 293, 294, 295, 296, 299, 300, 302, 304, 305, 306, 307, 308, 309, 310, 311, 313, 315, 316, 317, 319, 320, 321, 322, 323, 324, 326, 327, 328, 329, 330, 331, 334, 335, 336, 338, 339, 340, 342, 343, 344, 347, 348, 349, 350, 352, 354, 355, 356, 357, 358, 360, 361, 362, 364, 366, 367, 368, 369, 370, 371, 372, 373, 374, 377, 378, 379, 381, 383, 384, 385, 387, 389, 390, 391, 393, 394, 395, 396, 397, 398, 399, 400, 401, 402, 403, 407, 408, 410, 413, 415, 416, 417, 418, 419, 420, 421, 423, 424, 425, 427, 429, 430, 432, 434, 436, 437, 438, 439, 440, 441, 442, 443, 444, 445, 446, 447, 448, 452, 453, 454, 455, 456, 458, 459, 460, 461, 462, 465, 466, 468, 469, 470, 471, 472, 473, 474, 475, 476, 477, 479, 480, 482, 483, 484, 485, 486, 488, 489, 490, 491, 493, 494, 495, 496, 497, 498, 499, 500, 502, 503, 505, 506, 507, 508, 509, 510, 511, 512, 514, 515, 516, 518, 519, 521, 522, 523, 524, 526, 528, 529, 531, 533, 534, 535, 536, 537, 538, 539, 540, 541, 542, 543, 544, 545, 548, 549, 551, 552, 553, 555, 556, 557, 558, 560, 562, 564, 565, 566, 567, 568, 569, 570, 571, 572, 575, 576, 577, 578, 579, 580, 581, 582, 583, 584, 586, 587, 588, 589, 590, 591, 592, 593, 594, 595, 596, 597, 598, 599, 600, 601, 602, 603, 605, 606, 607, 609, 610, 611, 612, 614, 615, 616, 617, 618, 619, 620, 621, 624, 625, 627, 628, 629, 630, 631, 632, 634, 636, 637, 638, 639, 640, 641, 642, 643, 644, 645, 646, 649, 650, 651, 652, 653, 654, 655, 656, 657, 658, 660, 661, 662, 663, 666, 667, 670, 671, 672, 673, 674, 675, 676, 677, 679, 680, 682, 683, 684, 685, 686, 687, 689, 690, 691, 692, 693, 694, 696, 697, 698, 699, 700, 701, 702, 705, 707, 708, 709, 710, 711, 713, 714, 715, 716, 717, 718, 721, 722, 723, 724, 727, 728, 729, 732, 733, 734, 735, 737, 738, 739, 740, 741, 742, 743, 745, 746, 747, 749, 750, 752, 754, 755, 756, 757, 758, 760, 763, 764, 765, 767, 769, 770, 771, 772, 773, 774, 775, 777, 780, 781, 782, 783, 784, 785, 786, 787, 788, 789, 790, 791, 792, 793, 794, 795, 796, 797, 800, 801, 802, 803, 805, 806, 807, 808, 810, 812, 813, 814, 815, 816, 817, 818, 819, 820, 822, 823, 824, 825, 828, 830, 831, 832, 833, 834, 835, 836, 837, 838, 839, 841, 842, 843, 844, 845, 846, 847, 848, 852, 854, 856, 857, 858, 859, 860, 861, 862, 864, 865, 866, 867, 868, 869, 871, 872, 873, 875, 876, 877, 878, 879, 880, 882, 883, 885, 886, 888, 889, 890, 891, 893, 897, 898, 899, 900, 901, 903, 904, 905, 906, 907, 908, 909, 910, 911, 912, 913, 914, 915, 917, 918, 920, 921, 922, 923, 924, 925, 926, 928, 929, 930, 931, 932, 933, 934, 935, 936, 937, 938, 939, 940, 942, 943, 945, 946, 947, 950, 951, 952, 953, 954, 955, 958, 959, 960, 962, 963, 964, 965, 967, 968, 969, 970, 971, 973, 974, 975, 976, 977, 978, 979, 980, 981, 983, 984, 985, 986, 987, 988, 989, 991, 992, 994, 995, 996, 997, 998, 999, 1001, 1002, 1003, 1005, 1006, 1007, 1008, 1009, 1010, 1011, 1012, 1013, 1014, 1016, 1018, 1020, 1021, 1024, 1026, 1027, 1028, 1029, 1031, 1032, 1033, 1034, 1035, 1037, 1039, 1040, 1041, 1043, 1044, 1045, 1046, 1047, 1048, 1049, 1050, 1051, 1052, 1053, 1057, 1058, 1059, 1060, 1061, 1062, 1064, 1065, 1066, 1068, 1069, 1070, 1071, 1072, 1074, 1076, 1077, 1079, 1080, 1081, 1082, 1084, 1085, 1086, 1087, 1088, 1089, 1091, 1092, 1093, 1094, 1095, 1096, 1097, 1098, 1099, 1100, 1101, 1102, 1103, 1104, 1105, 1106, 1109, 1110, 1111, 1112, 1113, 1114, 1115, 1116, 1117, 1118, 1121, 1122, 1123, 1125, 1126, 1127, 1129, 1130, 1131, 1133, 1135, 1136, 1137, 1139, 1141, 1143, 1145, 1147, 1148, 1149, 1150, 1151, 1152, 1153, 1154, 1155, 1157, 1158, 1159, 1160, 1161, 1163, 1165, 1166, 1168, 1170, 1173, 1174, 1175, 1176, 1177, 1179, 1180, 1181, 1182, 1183, 1184, 1185, 1187, 1189, 1190, 1191, 1192, 1193, 1194, 1195, 1196, 1197, 1198, 1199, 1200, 1201, 1203, 1204, 1206, 1207, 1208, 1209, 1210, 1211, 1213, 1215, 1216, 1218, 1220, 1222, 1224, 1225, 1227, 1228, 1229, 1230, 1231, 1232, 1234, 1235, 1236, 1237, 1238, 1239, 1241, 1242, 1243, 1244, 1245, 1246, 1247, 1248, 1249, 1251, 1252, 1253, 1254, 1255, 1256, 1257, 1258, 1259, 1260, 1261, 1262, 1263, 1264, 1265, 1266, 1268, 1270, 1271, 1272, 1273, 1275, 1276, 1279, 1280, 1281, 1282, 1283, 1284, 1285, 1286, 1287, 1288, 1289, 1290, 1291, 1292, 1293, 1294, 1295, 1296, 1297, 1298, 1299, 1300, 1301, 1303, 1304, 1305, 1307, 1308, 1309, 1311, 1312, 1313, 1314, 1316, 1317, 1318, 1319, 1321, 1322, 1323, 1324, 1325, 1326, 1327, 1328, 1329, 1330, 1331, 1332, 1335, 1336, 1337, 1339, 1340, 1341, 1342, 1343, 1344, 1345, 1346, 1348, 1349, 1350, 1352, 1354, 1355, 1356, 1357, 1358, 1359, 1360, 1361, 1363, 1364, 1365, 1367, 1368, 1369, 1371, 1372, 1373, 1374, 1375, 1376, 1378, 1380, 1381, 1382, 1383, 1385, 1386, 1388, 1389, 1392, 1393, 1394, 1396, 1398, 1399, 1400, 1401, 1403, 1404, 1406, 1407, 1408, 1409, 1410, 1411, 1412, 1413, 1414, 1415, 1416, 1417, 1418, 1419, 1420, 1421, 1422, 1423, 1424, 1427, 1428, 1429, 1430, 1431, 1432, 1433, 1435, 1436, 1440, 1441, 1443, 1444, 1445, 1446, 1447, 1448, 1449, 1451, 1452, 1453, 1456, 1457, 1458, 1459, 1460, 1461, 1462, 1463, 1464, 1465, 1466, 1467, 1468, 1470, 1471, 1472, 1473, 1475, 1476, 1477, 1478, 1479, 1481, 1482, 1483, 1484, 1486, 1487, 1488, 1490, 1491, 1493, 1494, 1495, 1497, 1498, 1499, 1500, 1501, 1502, 1503, 1504, 1507, 1508, 1509, 1510, 1511, 1513, 1515, 1517, 1518, 1519, 1521, 1524, 1526, 1527, 1528, 1530, 1531, 1532, 1533, 1534, 1535, 1536, 1537, 1538, 1540, 1542, 1543, 1544, 1545, 1546, 1547, 1549, 1550, 1551, 1552, 1553, 1554, 1555, 1556, 1557, 1558, 1559, 1560, 1561, 1562, 1564, 1567, 1568, 1569, 1571, 1572, 1573, 1574, 1575, 1576, 1577, 1578, 1580, 1581, 1582, 1583, 1584, 1585, 1586, 1587, 1588, 1589, 1590, 1591, 1592, 1593, 1594, 1595, 1596, 1599, 1600, 1602, 1603, 1604, 1605, 1606, 1608, 1610, 1613, 1614, 1615, 1616, 1617, 1618, 1619, 1620, 1621, 1622, 1623, 1624, 1625, 1626, 1627, 1628, 1629, 1630, 1631, 1633, 1634, 1635, 1636, 1637, 1638, 1640, 1641, 1642, 1643, 1644, 1645, 1646, 1647, 1650, 1651, 1652, 1653, 1654, 1656, 1657, 1658, 1659, 1660, 1662, 1663, 1664, 1665, 1666, 1667, 1668, 1669, 1670, 1671, 1672, 1674, 1675, 1676, 1677, 1678, 1679, 1680, 1681, 1682, 1683, 1684, 1685, 1686, 1687, 1688, 1689, 1690, 1691, 1692, 1694, 1696, 1697, 1698, 1701, 1703, 1705, 1706, 1707, 1709, 1710, 1711, 1712, 1713, 1714, 1716, 1717, 1718, 1719, 1720, 1721, 1722, 1723, 1724, 1725, 1726, 1727, 1728, 1729, 1730, 1731, 1732, 1733, 1734, 1735, 1737, 1738, 1739, 1740, 1742, 1743, 1744, 1745, 1746, 1747, 1751, 1752, 1753, 1754, 1755, 1756, 1757, 1758, 1759, 1760, 1761, 1762, 1763, 1764, 1765, 1766, 1767, 1768, 1769, 1771, 1773, 1774, 1777, 1779, 1781, 1784, 1787, 1788, 1789, 1790, 1791, 1792, 1794, 1795, 1796, 1797, 1798, 1799, 1800, 1801, 1802, 1803, 1804, 1806, 1807, 1808, 1810, 1811, 1812, 1813, 1814, 1815, 1818, 1819, 1821, 1822, 1823, 1824, 1825, 1826, 1827, 1828, 1829, 1830, 1831, 1832, 1835, 1836, 1837, 1838, 1839, 1840, 1841, 1842, 1843, 1844, 1845, 1847, 1848, 1849, 1850, 1851, 1853, 1854, 1855, 1856, 1857, 1858, 1859, 1860, 1861, 1862, 1863, 1864, 1865, 1867, 1869, 1870, 1871, 1872, 1873, 1874, 1875, 1876, 1877, 1879, 1880, 1881, 1882, 1883, 1884, 1885, 1886, 1887, 1891, 1892, 1893, 1894, 1895, 1896, 1897, 1898, 1899, 1900, 1903, 1904, 1905, 1906, 1907, 1908, 1909, 1910, 1911, 1913, 1914, 1919, 1920, 1921, 1922, 1923, 1925, 1926, 1928, 1929, 1930, 1932, 1933, 1936, 1937, 1938, 1939, 1940, 1941, 1942, 1943, 1944, 1945, 1946, 1947, 1948, 1949, 1950, 1952, 1953, 1954, 1955, 1956, 1957, 1958, 1959, 1960, 1961], [351, 44, 194, 386, 547, 487, 659, 481, 359, 178, 79, 154, 527, 478, 573, 633, 467, 404, 550, 21, 435, 428, 426, 298, 504, 6, 301, 121, 281, 248, 207, 604, 520, 48, 412, 252, 38, 406, 195, 554, 405, 257, 74, 501, 152, 297, 622, 613, 17, 229, 110, 433, 532, 318, 411, 303, 341, 635, 382, 72, 513, 209, 585, 451, 365, 117, 574, 546, 346, 414, 392, 41, 375, 457, 247, 144, 626, 269, 186, 126, 332, 27, 563, 1, 184, 337, 167, 39, 530, 75, 409, 492, 245, 388, 517, 450, 137, 333, 561, 210, 2, 312, 132, 109, 95, 648, 623, 353, 76, 464, 314, 214, 525, 136, 463, 422, 431, 176, 647, 380, 256, 363, 287, 376, 240, 325, 283, 288, 608, 345, 559, 449, 1770, 1673, 1715, 874, 1217, 1496, 736, 688, 1036, 1563, 1030, 1315, 1019, 1523, 990, 972, 753, 1772, 1390, 821, 1233, 703, 853, 1833, 1142, 855, 1164, 1912, 1609, 1025, 1820, 1648, 1778, 1809, 730, 1162, 1522, 1384, 1749, 681, 1351, 1695, 1601, 961, 1132, 894, 948, 726, 799, 1171, 1775, 1107, 1334, 1708, 1310, 1267, 1816, 916, 1597, 1817, 1878, 804, 1489, 1119, 1962, 1485, 1442, 1347, 1700, 1927, 1167, 1454, 704, 1144, 849, 1140, 1015, 1598, 712, 1054, 1226, 1438, 1156, 720, 1202, 1450, 1178, 1124, 1931, 1924, 826, 768, 1455, 827, 927, 895, 725, 966, 1579, 1776, 850, 665, 1935, 759, 1277, 1649, 1607, 1915, 748, 1793, 1223, 668, 1120, 884, 1786, 1022, 1212, 956, 1785, 1320, 1565, 1750, 1426, 1890, 1078, 731, 896, 1205, 1916, 1566, 1868, 1055, 744, 1661, 1240, 1704, 1539, 851, 1083, 957, 1146, 1780, 1434, 1516, 1902, 1172, 1366, 778, 1090, 870, 1639, 1067, 1541, 902, 1250, 1075, 1889, 1306, 1693, 919, 1370, 798, 1611, 1063, 1888, 1221, 1782, 1570, 1402, 1134, 1302, 678, 1042, 1474, 1702, 811, 1529, 1278, 1169, 1387, 1377, 944, 1333, 941, 1783, 1934, 1741, 993, 1023, 1514, 1525, 1852, 1188, 1362, 1748, 1736, 695, 1951, 1397, 1918, 1437, 1214, 1901, 1834, 1405, 1269, 863, 1353, 1469, 1846, 1655, 809, 949, 762, 1548, 1219, 1492, 664, 751, 1274, 1512, 1338, 892, 1038, 1866, 1017, 1917, 779, 1699, 881, 1004, 1505, 829, 1379, 1000, 776, 719, 887, 761, 1056, 766, 1632, 1506, 1073, 1425, 706, 1128, 1439, 840, 1138, 669, 1612, 1520, 1108, 1480, 1395, 1186, 1391, 1805, 982])
([1, 2, 3, 4, 5, 6, 8, 11, 13, 15, 16, 17, 18, 21, 22, 23, 24, 25, 26, 27, 28, 29, 31, 32, 33, 34, 35, 38, 39, 41, 42, 43, 44, 45, 46, 48, 50, 51, 52, 53, 54, 55, 56, 57, 59, 60, 61, 62, 63, 64, 65, 68, 72, 74, 75, 76, 77, 79, 80, 81, 84, 86, 88, 89, 91, 92, 94, 95, 96, 97, 98, 100, 101, 102, 103, 104, 105, 107, 108, 109, 110, 113, 114, 115, 117, 118, 119, 121, 122, 123, 124, 126, 127, 128, 129, 131, 132, 133, 134, 135, 136, 137, 140, 141, 142, 143, 144, 145, 146, 150, 151, 152, 153, 154, 157, 158, 159, 161, 162, 163, 164, 165, 166, 167, 169, 170, 171, 172, 173, 174, 175, 176, 177, 178, 179, 181, 182, 184, 185, 186, 187, 188, 189, 190, 191, 193, 194, 195, 197, 198, 199, 200, 202, 203, 204, 206, 207, 208, 209, 210, 211, 212, 213, 214, 215, 217, 219, 221, 222, 223, 224, 225, 226, 227, 228, 229, 230, 233, 234, 235, 236, 237, 238, 239, 240, 242, 243, 245, 246, 247, 248, 251, 252, 254, 255, 256, 257, 258, 259, 260, 261, 262, 263, 265, 267, 268, 269, 270, 271, 273, 274, 275, 277, 278, 280, 281, 283, 284, 285, 286, 287, 288, 289, 290, 291, 294, 295, 296, 297, 298, 299, 300, 301, 302, 303, 305, 306, 307, 309, 311, 312, 313, 314, 315, 316, 318, 321, 323, 324, 325, 328, 329, 331, 332, 333, 334, 337, 338, 339, 340, 341, 343, 344, 345, 346, 347, 350, 351, 352, 353, 354, 355, 356, 359, 361, 362, 363, 364, 365, 366, 367, 368, 369, 370, 372, 373, 374, 375, 376, 377, 378, 379, 380, 381, 382, 383, 384, 385, 386, 387, 388, 389, 391, 392, 395, 396, 397, 398, 399, 400, 401, 402, 403, 404, 405, 406, 407, 408, 409, 410, 411, 412, 413, 414, 415, 416, 418, 419, 420, 422, 423, 424, 425, 426, 427, 428, 429, 430, 431, 432, 433, 434, 435, 436, 437, 439, 440, 444, 446, 447, 448, 449, 450, 451, 452, 454, 455, 456, 457, 458, 459, 461, 462, 463, 464, 465, 467, 468, 469, 470, 471, 472, 473, 474, 475, 476, 478, 480, 481, 483, 484, 485, 486, 487, 488, 490, 491, 492, 493, 494, 496, 498, 499, 500, 501, 502, 503, 504, 505, 506, 507, 508, 509, 510, 512, 513, 514, 515, 516, 517, 518, 519, 520, 522, 523, 524, 525, 527, 530, 532, 533, 534, 536, 539, 540, 541, 542, 543, 544, 545, 546, 547, 548, 549, 550, 551, 553, 554, 555, 556, 557, 559, 561, 562, 563, 564, 566, 567, 568, 569, 570, 571, 572, 573, 574, 575, 576, 577, 579, 581, 582, 583, 584, 585, 586, 587, 588, 589, 590, 591, 593, 594, 596, 597, 598, 600, 602, 603, 604, 605, 606, 607, 608, 609, 610, 611, 612, 613, 614, 616, 617, 618, 619, 620, 622, 623, 624, 625, 626, 627, 628, 631, 632, 633, 634, 635, 636, 637, 638, 639, 640, 641, 642, 643, 644, 645, 647, 648, 651, 652, 653, 654, 655, 656, 658, 659, 660, 661, 662, 663, 664, 665, 666, 667, 668, 669, 671, 672, 673, 674, 675, 676, 677, 678, 679, 680, 681, 682, 683, 685, 686, 687, 688, 689, 690, 691, 694, 695, 697, 698, 699, 700, 701, 702, 703, 704, 705, 706, 707, 709, 710, 712, 713, 716, 717, 718, 719, 720, 721, 722, 723, 724, 725, 726, 727, 730, 731, 733, 734, 735, 736, 738, 739, 740, 741, 742, 743, 744, 747, 748, 749, 750, 751, 752, 753, 754, 756, 757, 758, 759, 760, 761, 762, 763, 764, 765, 766, 767, 768, 769, 770, 771, 772, 773, 774, 775, 776, 777, 778, 779, 780, 781, 782, 783, 786, 787, 789, 790, 791, 792, 793, 796, 797, 798, 799, 800, 801, 802, 804, 805, 806, 809, 810, 811, 812, 813, 814, 815, 816, 817, 818, 819, 820, 821, 822, 823, 824, 825, 826, 827, 828, 829, 831, 832, 833, 834, 836, 837, 839, 840, 841, 842, 844, 845, 846, 847, 848, 849, 850, 851, 852, 853, 854, 855, 856, 857, 858, 860, 861, 862, 863, 865, 867, 868, 869, 870, 871, 872, 873, 874, 875, 876, 877, 879, 880, 881, 882, 883, 884, 885, 886, 887, 888, 890, 891, 892, 893, 894, 895, 896, 897, 898, 899, 901, 902, 903, 904, 905, 906, 907, 909, 910, 911, 913, 914, 915, 916, 917, 918, 919, 920, 922, 923, 925, 927, 929, 930, 931, 933, 934, 935, 936, 937, 939, 940, 941, 944, 946, 947, 948, 949, 950, 951, 952, 953, 955, 956, 957, 958, 959, 960, 961, 962, 963, 964, 965, 966, 970, 971, 972, 973, 974, 979, 980, 981, 982, 985, 986, 987, 988, 989, 990, 991, 992, 993, 995, 998, 999, 1000, 1001, 1003, 1004, 1005, 1006, 1007, 1008, 1009, 1010, 1013, 1014, 1015, 1016, 1017, 1018, 1019, 1020, 1022, 1023, 1024, 1025, 1026, 1028, 1030, 1032, 1033, 1034, 1035, 1036, 1037, 1038, 1039, 1040, 1041, 1042, 1043, 1044, 1045, 1046, 1047, 1048, 1049, 1050, 1051, 1052, 1053, 1054, 1055, 1056, 1057, 1058, 1059, 1060, 1061, 1062, 1063, 1064, 1065, 1066, 1067, 1068, 1069, 1071, 1073, 1074, 1075, 1076, 1078, 1079, 1080, 1081, 1083, 1085, 1087, 1088, 1089, 1090, 1092, 1093, 1094, 1096, 1097, 1099, 1100, 1101, 1102, 1103, 1104, 1105, 1106, 1107, 1108, 1109, 1110, 1111, 1112, 1114, 1115, 1116, 1117, 1118, 1119, 1120, 1122, 1123, 1124, 1126, 1128, 1129, 1130, 1132, 1133, 1134, 1135, 1136, 1137, 1138, 1139, 1140, 1141, 1142, 1143, 1144, 1146, 1148, 1150, 1152, 1153, 1155, 1156, 1157, 1158, 1159, 1162, 1163, 1164, 1165, 1166, 1167, 1168, 1169, 1170, 1171, 1172, 1173, 1174, 1176, 1177, 1178, 1179, 1180, 1181, 1182, 1183, 1184, 1185, 1186, 1187, 1188, 1190, 1191, 1193, 1194, 1195, 1196, 1197, 1198, 1199, 1200, 1201, 1202, 1203, 1204, 1205, 1208, 1209, 1210, 1212, 1213, 1214, 1216, 1217, 1218, 1219, 1221, 1222, 1223, 1224, 1225, 1226, 1227, 1230, 1233, 1234, 1235, 1236, 1237, 1238, 1239, 1240, 1241, 1242, 1243, 1244, 1245, 1247, 1250, 1251, 1252, 1253, 1254, 1255, 1256, 1257, 1258, 1259, 1260, 1261, 1262, 1263, 1264, 1266, 1267, 1268, 1269, 1272, 1274, 1275, 1277, 1278, 1280, 1281, 1282, 1283, 1284, 1285, 1286, 1287, 1288, 1289, 1290, 1291, 1296, 1297, 1298, 1299, 1300, 1302, 1303, 1304, 1306, 1307, 1308, 1309, 1310, 1312, 1313, 1314, 1315, 1316, 1317, 1318, 1319, 1320, 1321, 1322, 1324, 1325, 1327, 1328, 1331, 1333, 1334, 1337, 1338, 1339, 1340, 1341, 1342, 1343, 1344, 1347, 1348, 1349, 1350, 1351, 1352, 1353, 1354, 1355, 1356, 1358, 1360, 1362, 1363, 1365, 1366, 1370, 1372, 1374, 1377, 1378, 1379, 1382, 1383, 1384, 1385, 1387, 1389, 1390, 1391, 1392, 1394, 1395, 1397, 1398, 1399, 1400, 1401, 1402, 1403, 1404, 1405, 1406, 1407, 1408, 1409, 1411, 1412, 1415, 1417, 1418, 1419, 1421, 1423, 1424, 1425, 1426, 1427, 1428, 1429, 1430, 1434, 1435, 1437, 1438, 1439, 1440, 1442, 1443, 1445, 1446, 1447, 1448, 1450, 1451, 1453, 1454, 1455, 1456, 1457, 1458, 1460, 1462, 1463, 1464, 1465, 1466, 1467, 1468, 1469, 1470, 1471, 1472, 1473, 1474, 1475, 1476, 1477, 1479, 1480, 1481, 1482, 1483, 1484, 1485, 1487, 1489, 1490, 1491, 1492, 1493, 1495, 1496, 1497, 1498, 1500, 1501, 1502, 1503, 1504, 1505, 1506, 1508, 1509, 1512, 1514, 1515, 1516, 1517, 1518, 1519, 1520, 1521, 1522, 1523, 1524, 1525, 1526, 1529, 1530, 1531, 1532, 1533, 1539, 1540, 1541, 1542, 1544, 1545, 1546, 1548, 1549, 1550, 1552, 1554, 1555, 1556, 1557, 1559, 1562, 1563, 1564, 1565, 1566, 1567, 1568, 1569, 1570, 1571, 1572, 1576, 1577, 1578, 1579, 1580, 1583, 1585, 1586, 1587, 1589, 1591, 1592, 1594, 1595, 1596, 1597, 1598, 1599, 1601, 1602, 1605, 1607, 1608, 1609, 1610, 1611, 1612, 1613, 1616, 1617, 1618, 1620, 1621, 1622, 1623, 1624, 1626, 1627, 1629, 1630, 1631, 1632, 1633, 1634, 1635, 1637, 1638, 1639, 1641, 1644, 1645, 1647, 1648, 1649, 1650, 1651, 1652, 1654, 1655, 1656, 1657, 1658, 1659, 1661, 1662, 1664, 1665, 1666, 1667, 1668, 1669, 1670, 1671, 1672, 1673, 1674, 1675, 1677, 1678, 1679, 1681, 1682, 1683, 1684, 1685, 1687, 1688, 1690, 1691, 1692, 1693, 1694, 1695, 1697, 1698, 1699, 1700, 1702, 1703, 1704, 1705, 1706, 1707, 1708, 1709, 1710, 1711, 1712, 1713, 1714, 1715, 1718, 1720, 1721, 1722, 1723, 1724, 1725, 1727, 1728, 1729, 1730, 1732, 1733, 1734, 1735, 1736, 1739, 1741, 1742, 1743, 1744, 1746, 1747, 1748, 1749, 1750, 1751, 1752, 1754, 1755, 1757, 1758, 1759, 1761, 1763, 1765, 1766, 1767, 1770, 1771, 1772, 1773, 1774, 1775, 1776, 1777, 1778, 1780, 1781, 1782, 1783, 1785, 1786, 1787, 1788, 1789, 1790, 1792, 1793, 1794, 1795, 1796, 1797, 1798, 1799, 1800, 1801, 1802, 1803, 1804, 1805, 1806, 1807, 1809, 1810, 1811, 1812, 1813, 1814, 1816, 1817, 1818, 1819, 1820, 1821, 1822, 1823, 1824, 1825, 1826, 1827, 1828, 1829, 1830, 1831, 1832, 1833, 1834, 1835, 1836, 1837, 1838, 1841, 1842, 1843, 1844, 1845, 1846, 1847, 1848, 1850, 1852, 1853, 1854, 1855, 1856, 1857, 1859, 1860, 1861, 1862, 1865, 1866, 1867, 1868, 1869, 1871, 1873, 1874, 1875, 1876, 1878, 1879, 1880, 1883, 1884, 1885, 1886, 1887, 1888, 1889, 1890, 1892, 1893, 1894, 1895, 1896, 1897, 1898, 1899, 1900, 1901, 1902, 1903, 1904, 1905, 1906, 1907, 1908, 1909, 1910, 1911, 1912, 1913, 1914, 1915, 1916, 1917, 1918, 1919, 1920, 1923, 1924, 1925, 1926, 1927, 1929, 1931, 1933, 1934, 1935, 1936, 1938, 1941, 1942, 1945, 1946, 1947, 1949, 1950, 1951, 1953, 1954, 1955, 1957, 1959, 1960, 1961, 1962], [138, 349, 595, 116, 20, 69, 489, 47, 111, 482, 327, 168, 390, 83, 371, 558, 445, 342, 148, 630, 650, 438, 264, 615, 358, 552, 335, 649, 417, 336, 147, 30, 266, 393, 565, 272, 196, 308, 330, 149, 82, 49, 93, 646, 592, 421, 70, 244, 578, 497, 479, 460, 250, 85, 495, 466, 253, 537, 36, 87, 657, 521, 326, 357, 560, 180, 317, 19, 477, 276, 37, 320, 125, 66, 282, 453, 12, 293, 535, 531, 112, 601, 192, 139, 442, 529, 58, 526, 599, 348, 304, 310, 232, 160, 7, 183, 155, 71, 120, 360, 9, 156, 90, 443, 78, 279, 528, 99, 249, 220, 201, 511, 67, 10, 319, 538, 322, 231, 40, 629, 394, 292, 130, 106, 441, 241, 205, 218, 14, 73, 580, 621, 216, 1270, 1619, 1593, 1499, 1956, 1791, 1740, 1653, 746, 1839, 1877, 737, 912, 1211, 1369, 1332, 945, 1560, 1882, 1864, 1547, 1551, 1891, 1294, 1373, 1396, 1745, 1215, 1207, 968, 1336, 788, 1292, 1070, 859, 1939, 1444, 803, 1279, 1726, 1937, 684, 1410, 1077, 1581, 1206, 1851, 1295, 1461, 1756, 1027, 1940, 938, 1371, 1246, 1192, 1305, 1606, 1393, 1231, 997, 1121, 728, 1381, 1553, 1636, 1689, 1151, 1276, 1762, 838, 1588, 1273, 1717, 1293, 1160, 1375, 1952, 1511, 984, 1731, 1528, 1486, 1574, 835, 696, 866, 1696, 1881, 1029, 1380, 1536, 977, 1646, 1228, 1232, 1686, 785, 1346, 1625, 967, 708, 1357, 921, 843, 1095, 1220, 1229, 889, 1125, 1011, 1359, 1488, 978, 1082, 1330, 1301, 808, 1922, 1189, 1808, 1113, 1719, 1127, 1858, 1388, 1676, 1478, 932, 1764, 1815, 1872, 1928, 1361, 1642, 1510, 1701, 1441, 900, 1538, 1091, 1459, 1248, 1660, 1431, 1323, 1600, 1002, 996, 1932, 1175, 942, 1271, 1582, 715, 1535, 807, 794, 954, 1364, 830, 1768, 1452, 994, 1716, 1590, 711, 755, 1680, 1413, 1604, 1416, 1507, 969, 1784, 1367, 1414, 729, 1031, 1086, 1615, 1737, 1921, 784, 693, 1561, 1161, 1614, 1422, 1944, 1769, 1958, 928, 1311, 1537, 1131, 1265, 1943, 1863, 1543, 1663, 878, 1849, 1575, 1930, 1628, 1948, 1012, 1573, 714, 1603, 1386, 1753, 1779, 976, 1149, 1098, 732, 1534, 1021, 1368, 1640, 1558, 1147, 1432, 1760, 1420, 1494, 1449, 1329, 1154, 692, 1870, 1584, 1084, 926, 1643, 1738, 1436, 1840, 1335, 1527, 1513, 908, 1345, 943, 745, 983, 864, 1326, 1249, 1145, 924, 1376, 1072, 795, 1433, 670, 975])
([1, 2, 4, 5, 6, 7, 8, 9, 10, 12, 13, 14, 15, 16, 17, 19, 20, 21, 22, 26, 27, 28, 30, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 51, 52, 53, 54, 55, 56, 57, 58, 59, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 93, 94, 95, 96, 99, 101, 104, 106, 108, 109, 110, 111, 112, 115, 116, 117, 118, 119, 120, 121, 123, 124, 125, 126, 128, 130, 132, 133, 134, 135, 136, 137, 138, 139, 142, 143, 144, 145, 147, 148, 149, 150, 152, 154, 155, 156, 157, 158, 160, 161, 162, 163, 167, 168, 169, 171, 172, 173, 174, 175, 176, 177, 178, 179, 180, 181, 183, 184, 185, 186, 187, 188, 192, 194, 195, 196, 197, 198, 199, 201, 202, 203, 204, 205, 206, 207, 208, 209, 210, 211, 214, 215, 216, 217, 218, 219, 220, 221, 222, 225, 226, 228, 229, 230, 231, 232, 233, 235, 238, 239, 240, 241, 242, 243, 244, 245, 247, 248, 249, 250, 252, 253, 254, 255, 256, 257, 258, 260, 263, 264, 265, 266, 268, 269, 272, 273, 276, 277, 279, 281, 282, 283, 285, 286, 287, 288, 290, 292, 293, 294, 297, 298, 301, 303, 304, 305, 306, 308, 309, 310, 312, 313, 314, 315, 316, 317, 318, 319, 320, 321, 322, 323, 324, 325, 326, 327, 329, 330, 332, 333, 334, 335, 336, 337, 338, 339, 340, 341, 342, 343, 344, 345, 346, 347, 348, 349, 350, 351, 352, 353, 354, 356, 357, 358, 359, 360, 361, 362, 363, 364, 365, 366, 368, 369, 371, 373, 374, 375, 376, 377, 379, 380, 381, 382, 383, 385, 386, 387, 388, 389, 390, 391, 392, 393, 394, 395, 397, 398, 399, 400, 401, 402, 403, 404, 405, 406, 407, 408, 409, 410, 411, 412, 414, 415, 417, 418, 419, 420, 421, 422, 423, 424, 425, 426, 427, 428, 429, 430, 431, 432, 433, 435, 436, 437, 438, 439, 441, 442, 443, 444, 445, 447, 448, 449, 450, 451, 453, 455, 457, 458, 459, 460, 461, 462, 463, 464, 465, 466, 467, 470, 471, 472, 475, 477, 478, 479, 480, 481, 482, 483, 485, 486, 487, 489, 490, 491, 492, 493, 494, 495, 497, 498, 500, 501, 502, 503, 504, 505, 507, 508, 509, 510, 511, 512, 513, 516, 517, 518, 520, 521, 522, 523, 524, 525, 526, 527, 528, 529, 530, 531, 532, 533, 534, 535, 537, 538, 540, 541, 542, 543, 544, 545, 546, 547, 548, 550, 552, 553, 554, 558, 559, 560, 561, 563, 564, 565, 566, 568, 570, 572, 573, 574, 575, 576, 578, 580, 582, 583, 584, 585, 586, 588, 590, 592, 593, 594, 595, 596, 598, 599, 600, 601, 602, 603, 604, 606, 607, 608, 609, 610, 611, 612, 613, 614, 615, 616, 617, 618, 620, 621, 622, 623, 624, 625, 626, 627, 629, 630, 632, 633, 635, 636, 637, 640, 642, 643, 644, 645, 646, 647, 648, 649, 650, 652, 653, 654, 655, 656, 657, 658, 659, 661, 662, 663, 664, 665, 666, 667, 668, 669, 670, 671, 672, 674, 675, 676, 677, 678, 679, 681, 683, 684, 685, 686, 687, 688, 689, 690, 691, 692, 693, 694, 695, 696, 697, 698, 699, 700, 701, 702, 703, 704, 705, 706, 707, 708, 709, 710, 711, 712, 714, 715, 716, 719, 720, 721, 722, 723, 724, 725, 726, 727, 728, 729, 730, 731, 732, 736, 737, 738, 739, 740, 741, 742, 743, 744, 745, 746, 748, 749, 750, 751, 753, 755, 757, 758, 759, 761, 762, 763, 766, 767, 768, 769, 770, 771, 772, 773, 775, 776, 777, 778, 779, 781, 783, 784, 785, 786, 788, 791, 792, 794, 795, 796, 798, 799, 801, 802, 803, 804, 805, 807, 808, 809, 810, 811, 812, 814, 815, 817, 818, 819, 821, 822, 823, 826, 827, 828, 829, 830, 831, 833, 835, 837, 838, 840, 841, 843, 845, 849, 850, 851, 852, 853, 854, 855, 856, 857, 858, 859, 860, 862, 863, 864, 866, 870, 871, 872, 873, 874, 876, 877, 878, 879, 880, 881, 884, 885, 887, 888, 889, 890, 892, 893, 894, 895, 896, 897, 899, 900, 901, 902, 903, 905, 906, 907, 908, 909, 911, 912, 914, 916, 917, 918, 919, 920, 921, 924, 925, 926, 927, 928, 930, 932, 934, 935, 938, 939, 941, 942, 943, 944, 945, 947, 948, 949, 950, 951, 952, 954, 956, 957, 958, 959, 960, 961, 964, 965, 966, 967, 968, 969, 970, 971, 972, 974, 975, 976, 977, 978, 980, 981, 982, 983, 984, 985, 987, 988, 989, 990, 991, 992, 993, 994, 995, 996, 997, 998, 999, 1000, 1001, 1002, 1003, 1004, 1005, 1006, 1007, 1009, 1010, 1011, 1012, 1015, 1017, 1018, 1019, 1021, 1022, 1023, 1024, 1025, 1026, 1027, 1029, 1030, 1031, 1032, 1034, 1036, 1038, 1040, 1041, 1042, 1043, 1044, 1045, 1047, 1050, 1052, 1054, 1055, 1056, 1057, 1058, 1059, 1060, 1062, 1063, 1065, 1067, 1068, 1070, 1072, 1073, 1074, 1075, 1076, 1077, 1078, 1079, 1080, 1081, 1082, 1083, 1084, 1086, 1088, 1090, 1091, 1092, 1093, 1095, 1096, 1098, 1099, 1102, 1103, 1105, 1107, 1108, 1109, 1111, 1113, 1115, 1116, 1117, 1118, 1119, 1120, 1121, 1124, 1125, 1127, 1128, 1129, 1131, 1132, 1134, 1135, 1137, 1138, 1140, 1142, 1144, 1145, 1146, 1147, 1148, 1149, 1151, 1153, 1154, 1155, 1156, 1157, 1158, 1160, 1161, 1162, 1163, 1164, 1165, 1166, 1167, 1168, 1169, 1170, 1171, 1172, 1173, 1174, 1175, 1176, 1177, 1178, 1181, 1182, 1183, 1184, 1186, 1188, 1189, 1191, 1192, 1194, 1195, 1198, 1200, 1201, 1202, 1203, 1204, 1205, 1206, 1207, 1208, 1209, 1211, 1212, 1214, 1215, 1216, 1217, 1219, 1220, 1221, 1223, 1224, 1226, 1228, 1229, 1231, 1232, 1233, 1234, 1236, 1237, 1240, 1241, 1243, 1244, 1246, 1247, 1248, 1249, 1250, 1251, 1252, 1253, 1255, 1258, 1260, 1261, 1262, 1263, 1264, 1265, 1267, 1268, 1269, 1270, 1271, 1273, 1274, 1276, 1277, 1278, 1279, 1280, 1282, 1283, 1285, 1286, 1287, 1288, 1290, 1291, 1292, 1293, 1294, 1295, 1297, 1298, 1299, 1300, 1301, 1302, 1304, 1305, 1306, 1310, 1311, 1312, 1313, 1314, 1315, 1316, 1317, 1319, 1320, 1321, 1322, 1323, 1325, 1326, 1327, 1329, 1330, 1331, 1332, 1333, 1334, 1335, 1336, 1337, 1338, 1339, 1340, 1342, 1343, 1345, 1346, 1347, 1348, 1350, 1351, 1352, 1353, 1354, 1356, 1357, 1358, 1359, 1360, 1361, 1362, 1363, 1364, 1365, 1366, 1367, 1368, 1369, 1370, 1371, 1372, 1373, 1374, 1375, 1376, 1377, 1379, 1380, 1381, 1383, 1384, 1385, 1386, 1387, 1388, 1389, 1390, 1391, 1392, 1393, 1394, 1395, 1396, 1397, 1399, 1400, 1402, 1403, 1404, 1405, 1406, 1407, 1408, 1409, 1410, 1411, 1413, 1414, 1415, 1416, 1417, 1418, 1419, 1420, 1421, 1422, 1423, 1425, 1426, 1427, 1428, 1429, 1430, 1431, 1432, 1433, 1434, 1436, 1437, 1438, 1439, 1440, 1441, 1442, 1443, 1444, 1446, 1447, 1449, 1450, 1451, 1452, 1453, 1454, 1455, 1456, 1457, 1459, 1460, 1461, 1462, 1465, 1466, 1467, 1469, 1470, 1472, 1473, 1474, 1475, 1477, 1478, 1480, 1481, 1482, 1483, 1485, 1486, 1487, 1488, 1489, 1490, 1491, 1492, 1493, 1494, 1495, 1496, 1497, 1499, 1500, 1501, 1502, 1503, 1505, 1506, 1507, 1508, 1509, 1510, 1511, 1512, 1513, 1514, 1515, 1516, 1518, 1519, 1520, 1521, 1522, 1523, 1524, 1525, 1527, 1528, 1529, 1530, 1531, 1532, 1533, 1534, 1535, 1536, 1537, 1538, 1539, 1540, 1541, 1542, 1543, 1546, 1547, 1548, 1550, 1551, 1552, 1553, 1554, 1555, 1556, 1557, 1558, 1560, 1561, 1562, 1563, 1564, 1565, 1566, 1567, 1568, 1570, 1571, 1573, 1574, 1575, 1577, 1578, 1579, 1581, 1582, 1584, 1585, 1586, 1588, 1590, 1592, 1593, 1594, 1595, 1597, 1598, 1599, 1600, 1601, 1602, 1603, 1604, 1605, 1606, 1607, 1609, 1611, 1612, 1613, 1614, 1615, 1619, 1620, 1621, 1622, 1625, 1626, 1627, 1628, 1631, 1632, 1633, 1634, 1635, 1636, 1637, 1638, 1639, 1640, 1641, 1642, 1643, 1644, 1645, 1646, 1647, 1648, 1649, 1650, 1651, 1652, 1653, 1654, 1655, 1656, 1659, 1660, 1661, 1662, 1663, 1664, 1665, 1668, 1670, 1671, 1672, 1673, 1675, 1676, 1677, 1678, 1680, 1681, 1682, 1685, 1686, 1687, 1688, 1689, 1690, 1691, 1692, 1693, 1694, 1695, 1696, 1698, 1699, 1700, 1701, 1702, 1703, 1704, 1706, 1707, 1708, 1709, 1710, 1711, 1712, 1715, 1716, 1717, 1718, 1719, 1722, 1723, 1725, 1726, 1730, 1731, 1732, 1734, 1735, 1736, 1737, 1738, 1739, 1740, 1741, 1743, 1744, 1745, 1747, 1748, 1749, 1750, 1751, 1753, 1754, 1755, 1756, 1757, 1759, 1760, 1761, 1762, 1763, 1764, 1768, 1769, 1770, 1772, 1773, 1775, 1776, 1778, 1779, 1780, 1782, 1783, 1784, 1785, 1786, 1787, 1788, 1789, 1791, 1792, 1793, 1794, 1795, 1796, 1797, 1798, 1799, 1800, 1801, 1802, 1803, 1804, 1805, 1806, 1807, 1808, 1809, 1811, 1812, 1813, 1814, 1815, 1816, 1817, 1818, 1820, 1821, 1822, 1823, 1825, 1827, 1828, 1829, 1831, 1832, 1833, 1834, 1835, 1836, 1837, 1839, 1840, 1842, 1843, 1844, 1846, 1847, 1848, 1849, 1850, 1851, 1852, 1853, 1854, 1855, 1857, 1858, 1859, 1860, 1861, 1862, 1863, 1864, 1865, 1866, 1868, 1870, 1872, 1874, 1875, 1877, 1878, 1879, 1880, 1881, 1882, 1883, 1884, 1885, 1886, 1887, 1888, 1889, 1890, 1891, 1892, 1893, 1894, 1895, 1896, 1897, 1898, 1899, 1900, 1901, 1902, 1903, 1905, 1906, 1907, 1909, 1911, 1912, 1913, 1914, 1915, 1916, 1917, 1918, 1919, 1920, 1921, 1922, 1923, 1924, 1925, 1926, 1927, 1928, 1930, 1931, 1932, 1933, 1934, 1935, 1937, 1938, 1939, 1940, 1941, 1942, 1943, 1944, 1945, 1947, 1948, 1951, 1952, 1955, 1956, 1957, 1958, 1962], [474, 141, 587, 224, 246, 605, 577, 234, 60, 191, 102, 641, 262, 555, 651, 514, 200, 378, 212, 328, 539, 33, 23, 473, 146, 446, 114, 413, 456, 25, 274, 295, 519, 581, 227, 32, 151, 300, 434, 289, 638, 193, 631, 660, 284, 355, 140, 571, 189, 166, 18, 29, 98, 440, 557, 454, 469, 499, 159, 127, 307, 452, 296, 113, 190, 271, 213, 549, 416, 484, 597, 103, 506, 251, 92, 515, 107, 164, 3, 468, 331, 105, 372, 619, 131, 270, 639, 259, 34, 129, 634, 81, 370, 261, 11, 496, 591, 476, 384, 165, 50, 628, 237, 97, 275, 100, 280, 153, 396, 223, 299, 367, 589, 488, 551, 122, 291, 311, 267, 556, 569, 24, 31, 536, 236, 567, 278, 170, 579, 182, 562, 302, 1257, 1746, 1150, 733, 1152, 1767, 1727, 718, 1318, 848, 1238, 1679, 682, 1424, 1674, 1016, 904, 1544, 1197, 953, 813, 1341, 1289, 1610, 754, 1869, 1179, 937, 1110, 680, 1908, 1946, 963, 1245, 1871, 1028, 1104, 1623, 1046, 825, 1445, 839, 1349, 1033, 1569, 1545, 800, 1112, 816, 1742, 986, 1781, 1100, 1398, 1227, 793, 1758, 1949, 946, 865, 1953, 1684, 1064, 1468, 915, 836, 1225, 1873, 875, 1720, 1401, 747, 1242, 673, 1187, 962, 1729, 1281, 1476, 1435, 1777, 1106, 1066, 1596, 1618, 1039, 1218, 883, 1199, 1517, 1774, 1666, 834, 752, 973, 868, 1254, 1355, 1658, 1222, 1013, 1838, 913, 790, 1307, 1235, 898, 1526, 1771, 1008, 1344, 1713, 1328, 756, 734, 1587, 1669, 1130, 1959, 1936, 844, 765, 1141, 940, 1126, 867, 1549, 1213, 1504, 1069, 824, 1266, 1589, 1185, 1094, 922, 1448, 789, 832, 1193, 820, 847, 861, 1087, 1159, 1910, 1766, 1929, 797, 1133, 1085, 764, 1624, 1733, 1324, 1378, 1143, 1498, 1608, 1617, 1309, 1765, 1576, 774, 869, 1484, 713, 1259, 1960, 1961, 882, 1284, 1819, 1667, 1458, 1180, 1630, 1867, 923, 1479, 1053, 735, 1464, 1014, 1683, 1752, 1196, 1061, 955, 891, 936, 806, 1463, 1724, 933, 1051, 1035, 1275, 1705, 1048, 1136, 1845, 1790, 1580, 846, 1114, 979, 1412, 1657, 1583, 1101, 886, 787, 1071, 842, 1841, 1591, 1616, 1190, 1089, 910, 1049, 1824, 1230, 1826, 1721, 1876, 1123, 1629, 1830, 782, 1382, 1954, 1037, 1272, 1020, 1714, 929, 1950, 1697, 1296, 1256, 1210, 1856, 931, 1728, 780, 1810, 1239, 1303, 1471, 1559, 1139, 1122, 717, 1572, 1308, 1904, 1097, 760])We will perform variable selection by adding variables one at a time, but we will constrain the model to start with BIO1.
variables!(model, ForwardSelection, folds; included=[1])☑️ PCATransform → Logistic → P(x) ≥ 0.508 🗺️The call to variables! will modify the model in two significant ways: it will select a subset of the variables, and then it will train the model using all the data available for training.
We can check that the model is trained:
istrained(model)trueWe can also inspect the list of selected variables:
variables(model)5-element Vector{Int64}:
1
2
8
13
15Index of variables
Models can be reset at any time, which means that dropped variables can be re-included. Therefore, the index of a variable in a model never refers to the list of included variables, but to the entire pool of variables that the model had access to when it was created.
At this point, we have a trained model, that is ready to be used.
Measuring the model performance
Now that we have identified the optimal set of variables, it is important to report the expected performance of this model. This is done internally when selecting the variables, but it is always a good idea to report on multiple measures of model performance, as they will indicate what type of predictions the model does well.
cval, ctrn = crossvalidate(model);This function returns a series of ConfusionMatrix vectors, which can be given as arguments to many measures of classifier performance. Different pages in the manual will use different approaches for validation, but the default that is enforced throughout the package is that Matthew's Correlation Coefficient is the best measure of performance. Whenever a function needs to decide on a "best" model, it will likely have a keyword to change this if necessary.
We can look at the first confusion matrix for the first split on the validation data:
cval[1](tp: 55, fp: 18; fn: 12, tn: 113)This gives the number of true/false positive/negatives for this specific split of the data.
PR and ROC curves
If you want to visualize these curves, there is a vignette on how to measure them.
The crossvalidate method may also take a second argument, which is a split of the dataset according to different criteria. The output of kfold we used earlier is one example, and the package offers several more.
To see how the model performs across a variety of measures, we will look at the averaged value of a few of them over the training and validation data.
ms = [mcc, tpr, fpr, fnr, tnr, ppv, npv]
M = permutedims([
m(c) for m in ms, c in [cval, ctrn]
]);
M = hcat(["Training", "Validation"], M);
pretty_table(
M;
alignment = [:l, :c, :c, :c, :c, :c, :c, :c],
backend = :markdown,
column_labels = ["Dataset"; uppercase.(string.(ms))],
formatters = [fmt__printf("%5.3f", [2, 3, 4, 5, 6, 7, 8])],
)| Dataset | MCC | TPR | FPR | FNR | TNR | PPV | NPV |
|---|---|---|---|---|---|---|---|
| Training | 0.691 | 0.834 | 0.131 | 0.166 | 0.869 | 0.768 | 0.911 |
| Validation | 0.707 | 0.849 | 0.127 | 0.151 | 0.873 | 0.773 | 0.919 |
This provides us with an estimate of the performance of our model, which will be useful to guide interpretation.
Prediction on the testing data
Back at the data collection step, we had also downloaded some testing data. We we will use them to make predictions on the environmental conditions associated to these observations. Note that we are indexing a vector of layers by a vector of occurrences, and this will return a matrix.
Xₜ = L[test_records];This matrix can be passed to the model (and the predict function) to generate the presence/absence prediction for this testing data. Note that the Xₜ matrix has the instances on the columns, which is more memory efficient and respect the convention of storing instances as vectors.
yₜ = predict(model, Xₜ);Dimension of input variables
Because the model keeps the variables that are not included, it must be given the full list of variables in order to make a prediction. In this case, even though we have selected variables, we are passing all 19 BioClim variables to make the prediction. This allows us to change the list of included variables at any point in the future.
We can calculate the accuracy of this prediction (which is appropriate to do here since the testing data are presence-only, so we do not really have class balance issues):
accuracy(yₜ, ones(Bool, length(yₜ)))0.872This indicates the proportion of presences that our model has correctly predicted. This model is doing a much better job at predicting the presences from the testing data than we should have expected based on the cross-validation results alone, which is great!
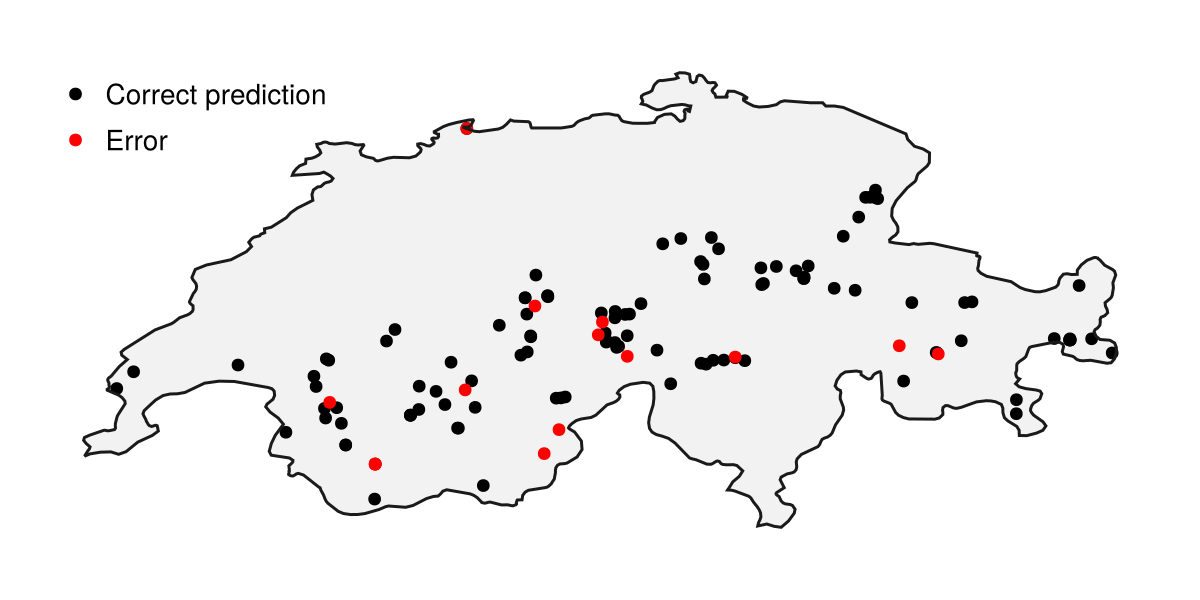
Code for the figure
f = Figure(; size = (600, 300))
ax = Axis(f[1, 1]; aspect = DataAspect())
poly!(ax, aoi; color = :grey95)
scatter!(ax, test_records[findall(yₜ)]; color = :black, label = "Correct prediction")
scatter!(ax, test_records[findall(!, yₜ)]; color = :red, label = "Error")
hidedecorations!(ax)
hidespines!(ax)
lines!(ax, aoi; color = :grey10)
axislegend(ax; position = :lt, framevisible = false)Understanding the model
Before moving on to making spatial predictions, it is important to understand what the model is doing to move from a value of the covariates to a prediction. For this, we can realy on functions that perform model explanations.
Model interpretability
Both model interpretability and feature importance have their own vignettes, which cover the essential options. There are additional techniques for model explanation that are not covered in this vignette.
We will estimate the importance of variables through the PartialDependence measure, which captures the variable that, on average across all training data, has the most different effect on prediction when its value is varied.
vimp = [featureimportance(PartialDependence, model, v) for v in variables(model)]
vimp ./= sum(vimp)
vord = sortperm(vimp, rev=true)
Dict(zip(layers(chelsa_bioclim)[variables(model)], vimp))Dict{String, Float64} with 5 entries:
"BIO8" => 0.224914
"BIO1" => 0.35245
"BIO13" => 0.186271
"BIO2" => 0.0790623
"BIO15" => 0.157303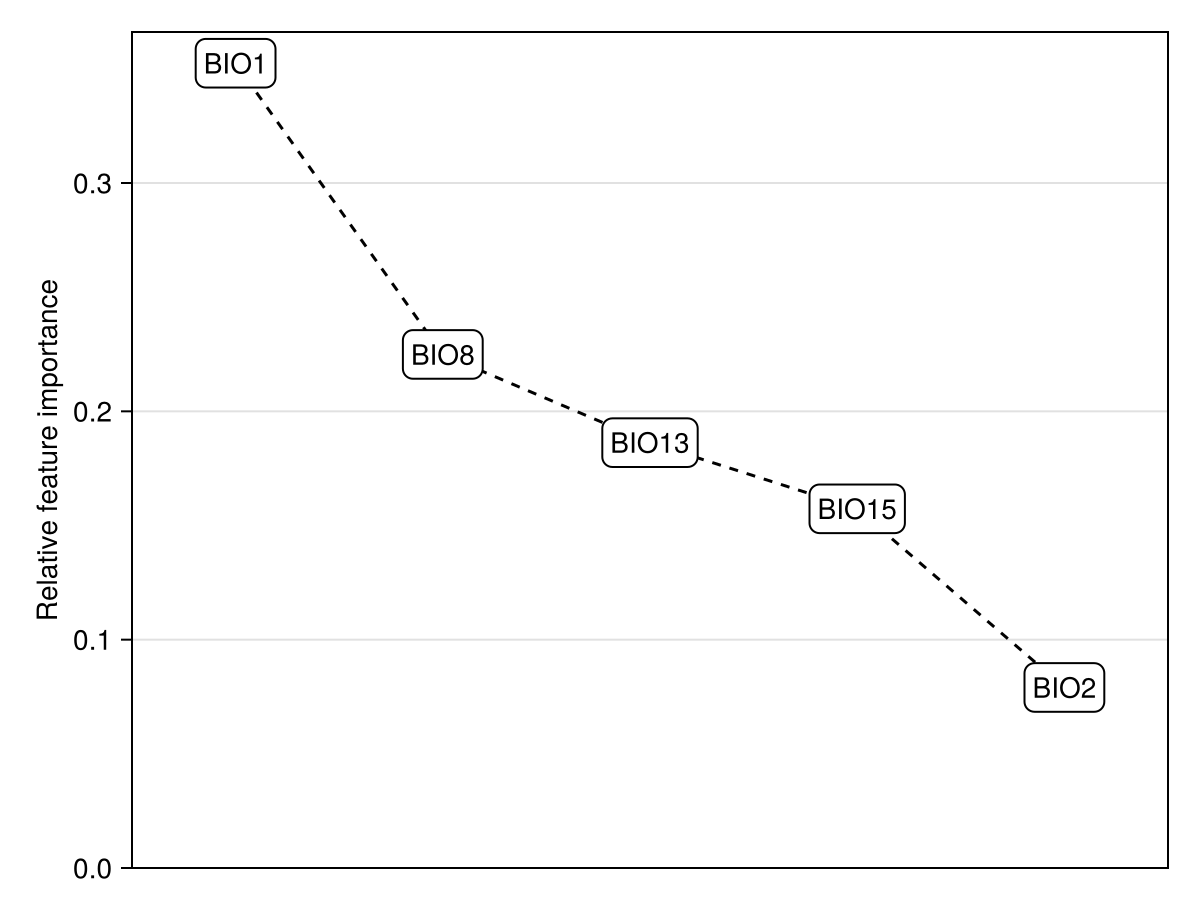
Code for the figure
f = Figure()
ax = Axis(f[1,1], ylabel="Relative feature importance")
lines!(ax, 1:length(vimp), vimp[vord], color=:black, linestyle=:dash)
textlabel!(ax, 1:length(vimp), vimp[vord], layers(chelsa_bioclim)[variables(model)[vord]])
ylims!(ax, low=0.0)
hidexdecorations!(ax)
xlims!(ax, 0.5, length(variables(model)) + 0.5)We will limit our analysis to the two most important variables:
v1, v2 = variables(model)[vord[1:2]]2-element Vector{Int64}:
1
8If we want to get more information about which these two are, we can use the layerdescriptions function to get plain-text information:
varnames = [
layerdescriptions(chelsa_bioclim)[l]
for l in layers(chelsa_bioclim)[[v1, v2]]
]2-element Vector{String}:
"Annual Mean Temperature"
"Mean Temperature of Wettest Quarter"To start with, we can look at the partial response of the model to the most important variables (the rugplots for the presences is at the bottom, and the one for the pseudo-absences at the top).
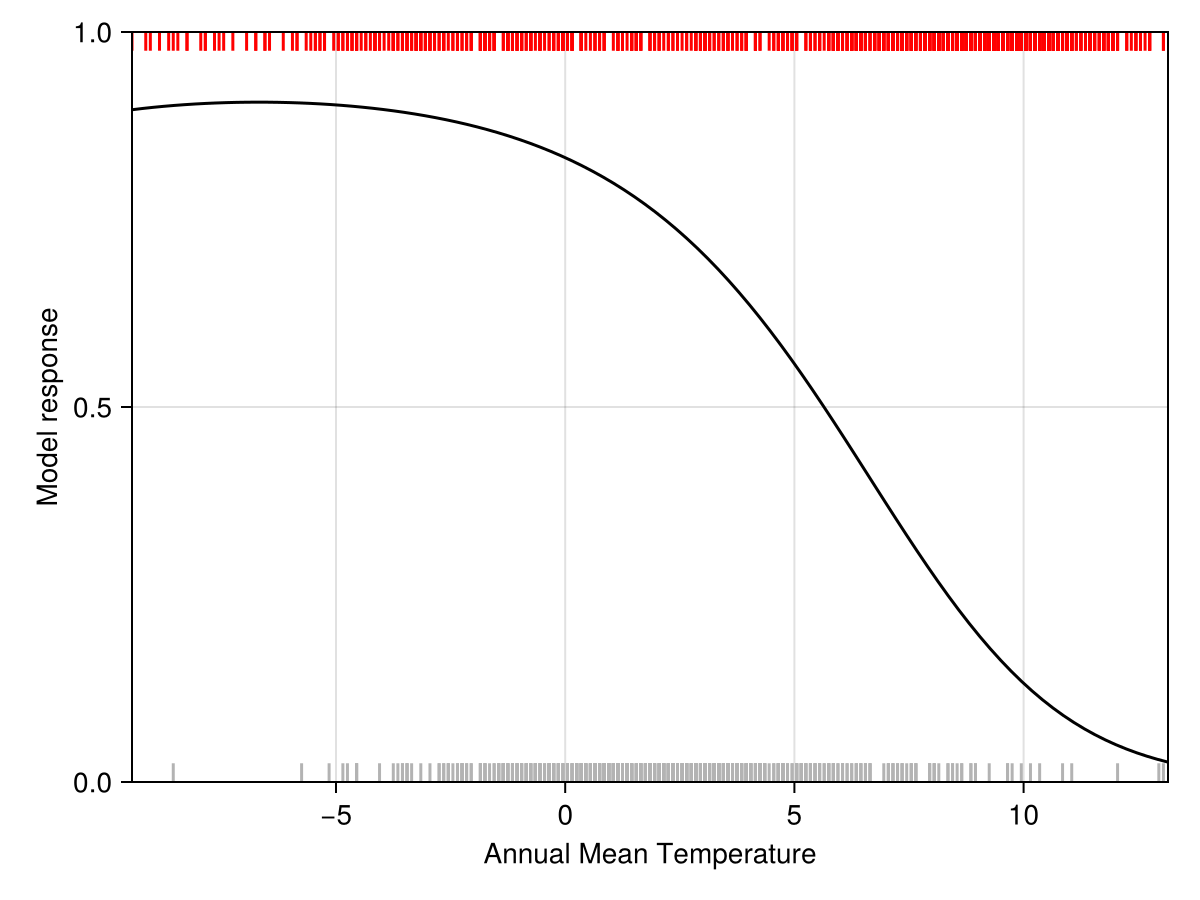
Code for the figure
f = Figure()
ax = Axis(f[1, 1]; xlabel=varnames[1], ylabel="Model response")
lines!(ax, explainmodel(PartialResponse, model, v1, 100; threshold=false)..., color=:black)
vlines!(ax, features(model, v1)[findall(labels(model))], ymax=0.025, color=:grey70)
vlines!(ax, features(model, v1)[findall(.!labels(model))], ymin=1-0.025, color=:red)
ylims!(ax, 0, 1)
tightlimits!(ax)We can also look at the combination of the two most important variables. This is not a statement about their importance as much as it is a statement about the fact that we are limited in the number of dimensions we can examine at once: a single variable is a line, two variables are a surface, and after this it gets complicated.
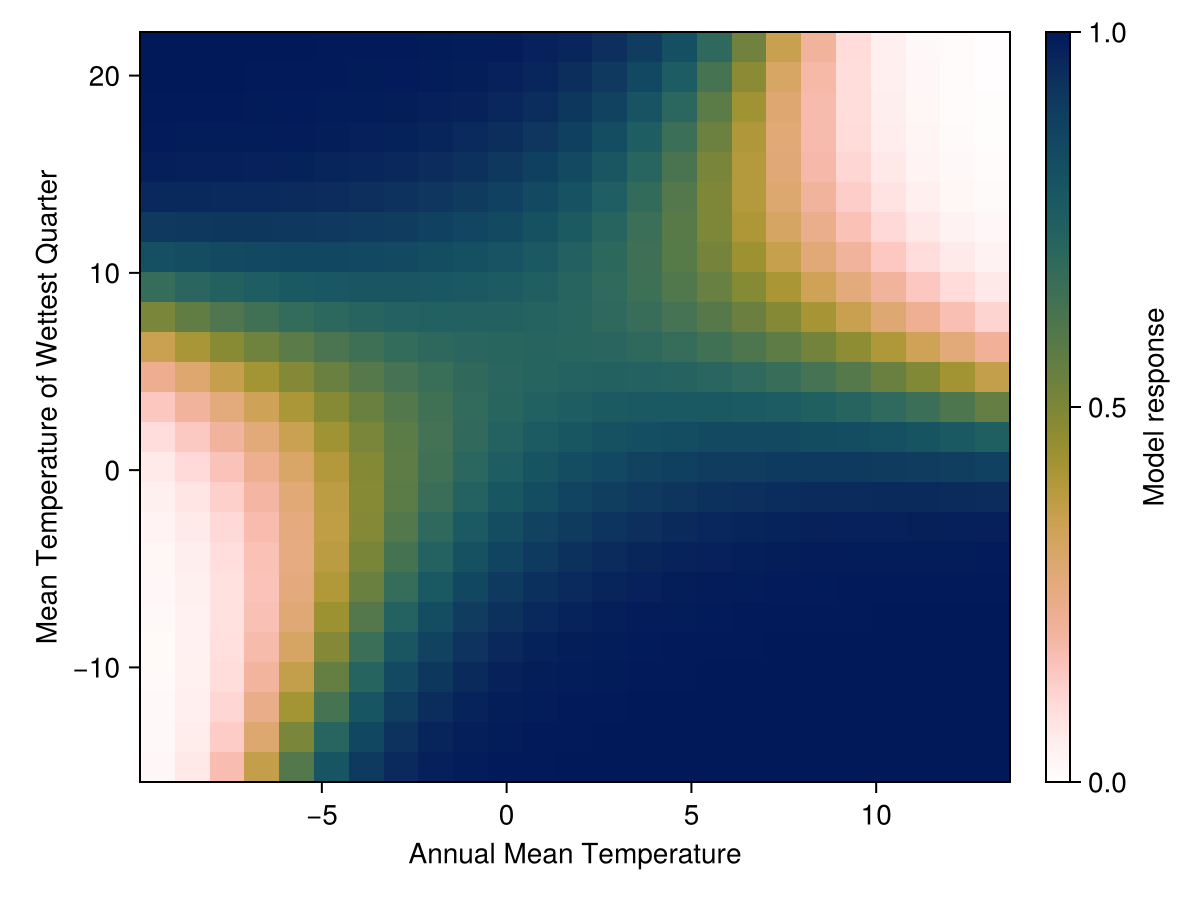
Code for the figure
f = Figure()
ax = Axis(f[1,1], xlabel=varnames[1], ylabel=varnames[2])
hm = heatmap!(ax,
explainmodel(PartialResponse, model, (v1, v2), 25; threshold=false)...,
colormap=Reverse(:batlowW), colorrange=(0, 1)
)
Colorbar(f[1,2], hm, label="Model response")Note that these responses do not provide the full picture of why the model makes a specific prediction. If we are interested in the effect of a variable on a specific prediction, the right tool is to use Shapley values. They link the value of a given variable to a departure from the average prediction.
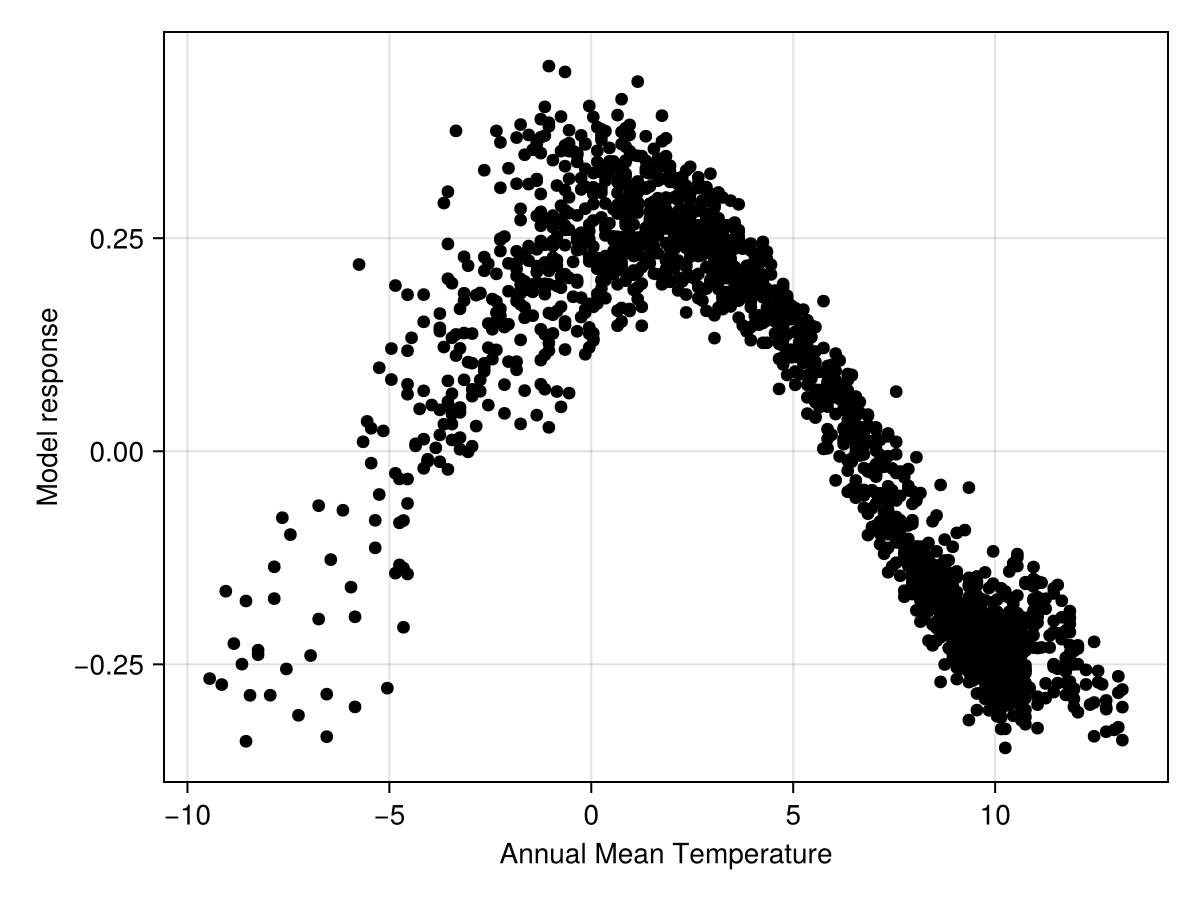
Code for the figure
f = Figure()
ax = Axis(f[1, 1]; xlabel=varnames[1], ylabel="Model response")
scatter!(ax, explainmodel(ShapleyMC, model, v1; threshold=false)..., color=:black)Shapley values are a little more informative, because they account for the value of all other variables at the point for which we ask for an explanation. For now, this is enough model explanation, and we will move on to making the first spatial prediction.
Spatial predictions
We can predict using the model by giving a vector of layers as an input. This will return the output as a layer as well:
P = predict(model, L; threshold = false)🗺️ A 239 × 543 layer with 70285 Float64 cells
Spatial Reference System: +proj=longlat +datum=WGS84 +no_defs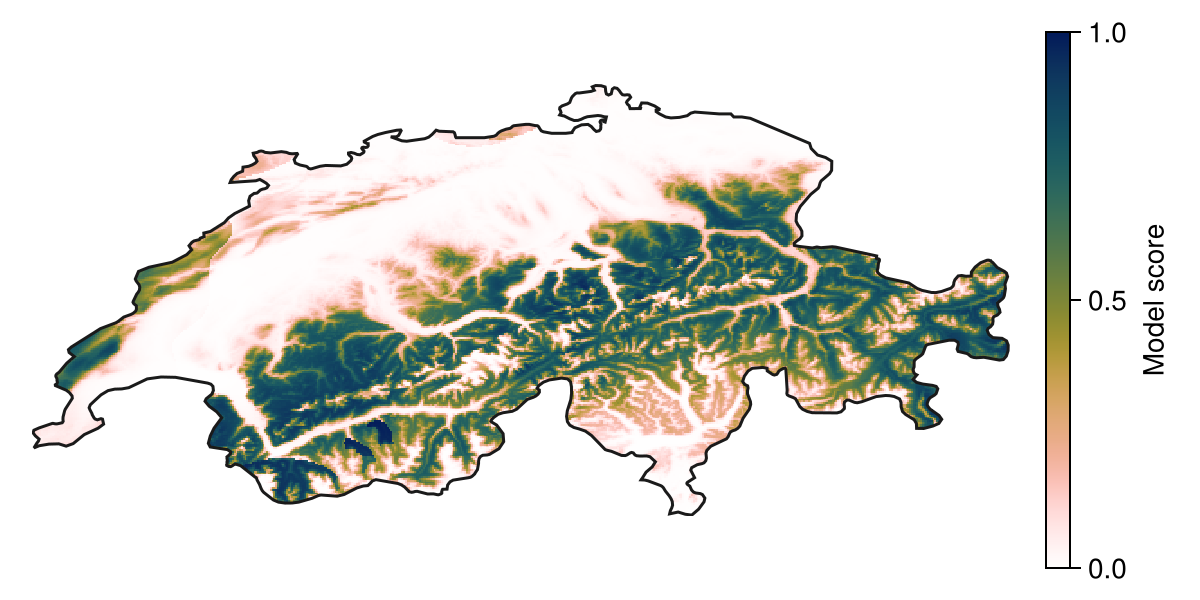
Code for the figure
f = Figure(; size = (600, 300))
ax = Axis(f[1, 1]; aspect = DataAspect())
hm = heatmap!(ax, P; colormap = Reverse(:batlowW), colorrange = (0, 1))
Colorbar(f[1, 2], hm, label="Model score")
lines!(ax, aoi; color = :grey10)
hidedecorations!(ax)
hidespines!(ax)Model score and probabilities
Whether this model score is an actual probability depends on whether the model is well calibrated. Refer to the vignette on calibration for more. Logistic regression tends to give pretty-well calibrated models.
We can look at the median suitability using zonal statistics for this species across the bioregions that are covering this territory, to identify the more favorable region:
bioregions = getpolygon(PolygonData(OneEarth, Bioregions))["Region" => "Western Eurasia"]
score_by_zone = byzone(
x -> isempty(x) ? NaN : median(x),
P,
bioregions,
"Name"
)
filter(v -> !isnan(v.second), score_by_zone)Dict{String, Float64} with 2 entries:
"Alps & Po Basin Mixed Forests" => 0.520262
"European Interior Mixed Forests" => 0.0168708In the above prediction, we have requested the model score, as opposed to the predicted presence. We can get the presence by not changing the threshold argument:
R = predict(model, L)🗺️ A 239 × 543 layer with 70285 Bool cells
Spatial Reference System: +proj=longlat +datum=WGS84 +no_defs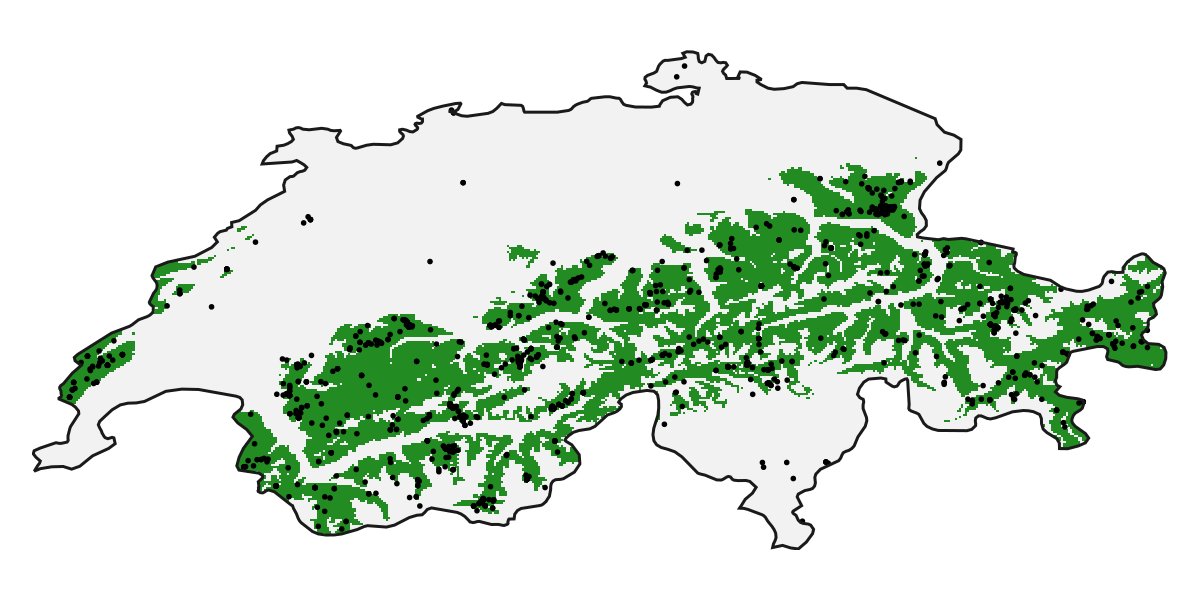
Code for the figure
f = Figure(; size = (600, 300))
ax = Axis(f[1, 1]; aspect = DataAspect())
poly!(ax, aoi; color = :grey95)
heatmap!(ax, R; colormap = [:transparent, :forestgreen])
scatter!(ax, records, color=:black, markersize=4)
lines!(ax, aoi; color = :grey10)
hidedecorations!(ax)
hidespines!(ax)Spatial explanations
When called with a vector of layers as additional arguments, the methods for model explanation will return a spatial version of the model response. For example, we can look at the effect of the most important variable on the prediction:
partial1 = explainmodel(PartialResponse, model, v1, L; threshold = false)🗺️ A 239 × 543 layer with 70285 Float64 cells
Spatial Reference System: +proj=longlat +datum=WGS84 +no_defs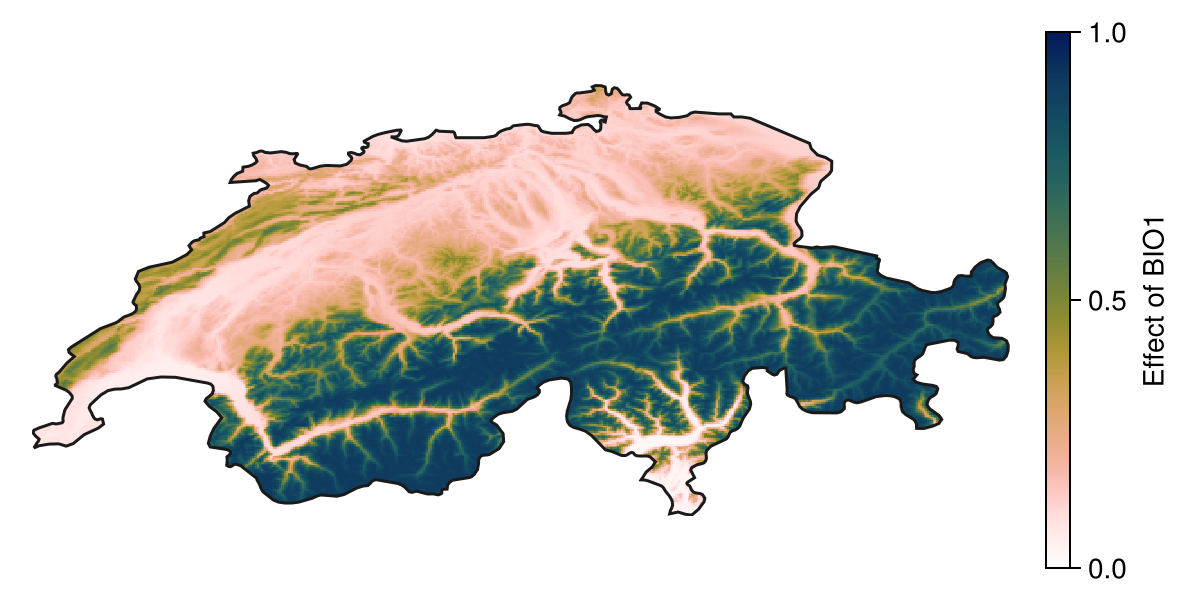
Code for the figure
f = Figure(; size = (600, 300))
ax = Axis(f[1, 1]; aspect = DataAspect())
hm = heatmap!(ax, partial1; colormap = Reverse(:batlowW), colorrange = (0, 1))
Colorbar(f[1, 2], hm, label="Effect of $(layers(chelsa_bioclim)[v1])")
lines!(ax, aoi; color = :grey10)
hidedecorations!(ax)
hidespines!(ax)This is particularly important to look at the local effect of the variables on each prediction, for example using Shapley values:
shapley1 = explainmodel(ShapleyMC, model, v1, L; threshold = false)🗺️ A 239 × 543 layer with 70285 Float64 cells
Spatial Reference System: +proj=longlat +datum=WGS84 +no_defsNote that the Shapley values are expressed as a deviation from the average prediction, and so positive values correspond to the presence class being more likely compared to the baseline.
Both the explain and partialresponse functions can accept keywords that are passed to predict - in this tutorial, we use threshold=false to provide an explanation on the score returned by the model, but we can also request an explanation of the binary response (this is usually less informative).
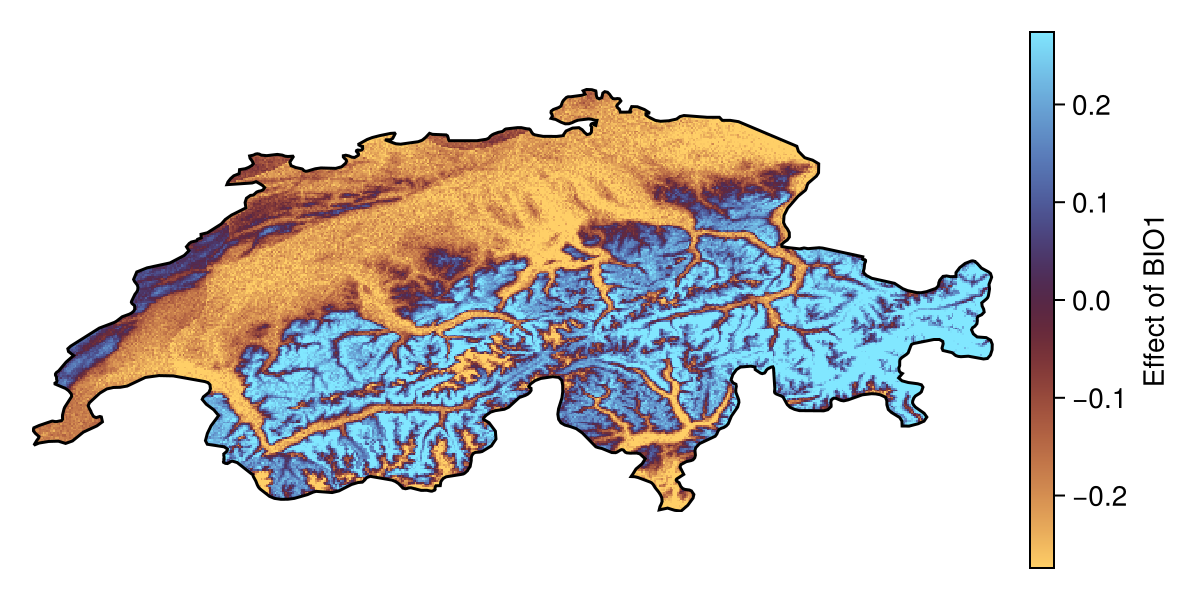
Code for the figure
col_lims = maximum(abs.(quantile(shapley1, [0.1, 0.9]))) .* (-1, 1)
f = Figure(; size = (600, 300))
ax = Axis(f[1, 1]; aspect = DataAspect())
hm = heatmap!(ax, shapley1; colormap = :managua, colorrange = col_lims)
Colorbar(f[1, 2], hm, label="Effect of BIO$(v1)")
lines!(ax, aoi; color = :black)
hidedecorations!(ax)
hidespines!(ax)We will now identify which of these variables is the most locally important to make the final prediction:
S = explainmodel(ShapleyMC, model, L; threshold = false)19-element Vector{SDMLayer{Float64}}:
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)
🗺️ A 239 × 543 layer (70285 Float64 cells)We can rely on the mosaic function to find the variable that has the largest absolute contribution to moving the prediction away from the average. This serves as a way to identify the most locally important variable:
keyvariable = mosaic(argmax, map(x -> abs.(x), S))🗺️ A 239 × 543 layer with 70285 Int64 cells
Spatial Reference System: +proj=longlat +datum=WGS84 +no_defsIt is important to notice that, even though our model relies on a PCA to transform the variables, we are looking at the importance of the variables before the transformation. We can, therefore, think biologically about the result of this analysis, regardless of the optimal way to project these raw covariates into latent variables for the classifier.
Because this too is returned as a layer, it can be mapped as well (after a little bit of mischief with the color palette):
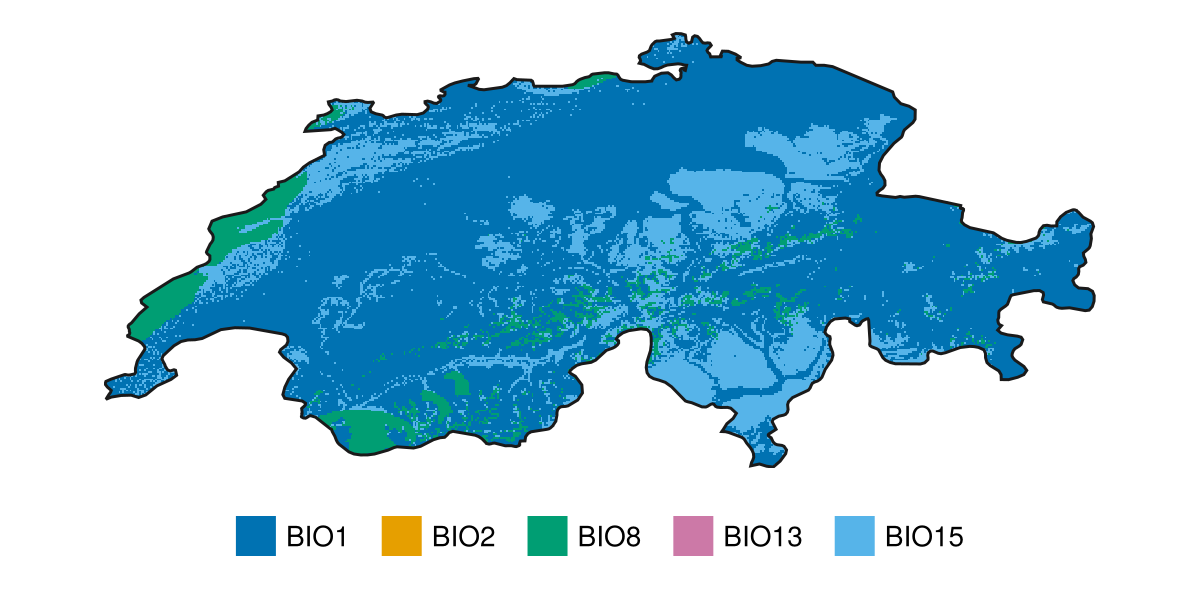
Code for the figure
f = Figure(; size = (600, 300))
colors = Makie.wong_colors()
n_var = length(variables(model))
n_col = length(colors)
if n_var > n_col
append!(colors, fill(colorant"#ececec", n_var - n_col))
end
palette = cgrad(colors[1:n_var], n_var, categorical=true)
ax = Axis(f[1, 1]; aspect = DataAspect())
heatmap!(ax, keyvariable; colormap = palette)
lines!(ax, aoi; color = :grey10)
hidedecorations!(ax)
hidespines!(ax)
Legend(
f[2, 1],
[PolyElement(; color = colors[i]) for i in 1:length(variables(model))],
["BIO$(b)" for b in variables(model)];
orientation = :horizontal,
nbanks = 1,
framevisible = false,
vertical = false,
)Bootstrap for uncertainty
As the last step, we will attempt to quantify the uncertainty of our model using non-parametric bootstrap. This is simply achieved by wrapping our model into a new constructor:
bootstrap = Bagging(model, 100)
train!(bootstrap){PCATransform → Logistic → P(x) ≥ 0.508} × 100All of these models have the same training data, but different training instances. We can measure the balanced out-of-bag error on this model:
outofbag(bootstrap) |> M -> 1 - balancedaccuracy(M)0.13271085659348125More importantly for this use-case, the bootstrapped model can make predictions using different information to aggregate the results of all models. A good measure of uncertainty is, for example, to use the inter-quartile range:
uncertainty = predict(bootstrap, L; threshold=false, consensus=iqr)🗺️ A 239 × 543 layer with 70285 Float64 cells
Spatial Reference System: +proj=longlat +datum=WGS84 +no_defsConformal prediction
The package offers functions for conformal prediction, and a more fully worked-out example of spatial projection of uncertainty. This is a more advanced feature, and for now we will be quite happy with good old non-parametric bootstrap. See also the vignette on covariate shift to measure how far away the prediction data are from the training data.
We can now map this uncertainty, to identify where the models in the bootstrapped ensemble disagree the most, and by how much they disagree:
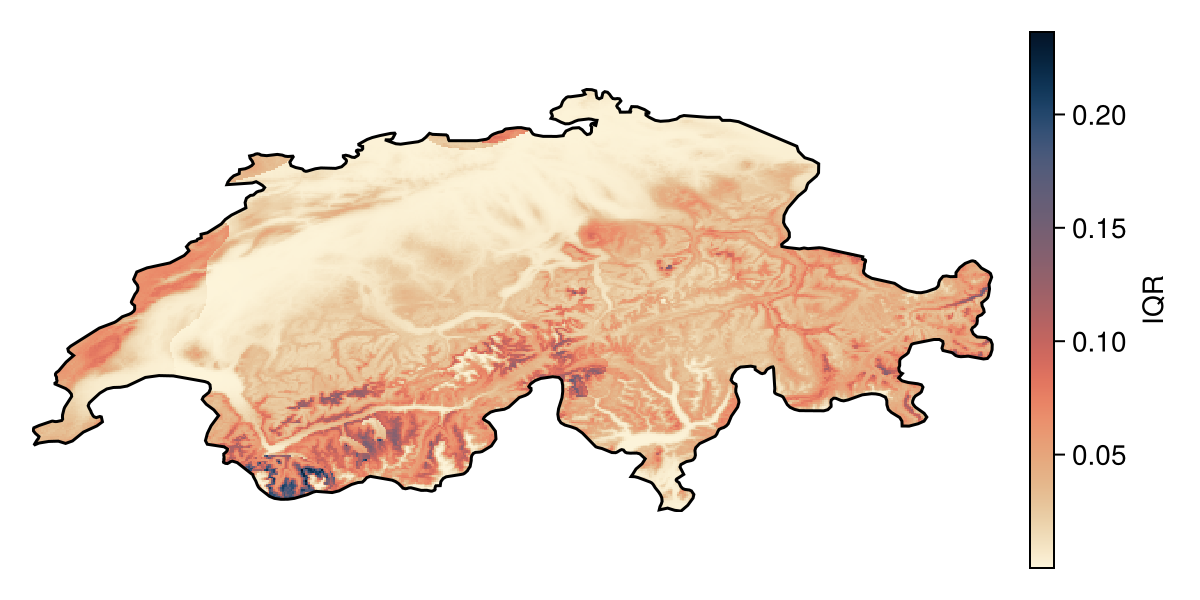
Code for the figure
f = Figure(; size = (600, 300))
ax = Axis(f[1, 1]; aspect = DataAspect())
hm = heatmap!(ax, uncertainty; colormap = Reverse(:lipari))
Colorbar(f[1, 2], hm, label="IQR")
lines!(ax, aoi; color = :black)
hidedecorations!(ax)
hidespines!(ax)This information can be jointly visualized with the prediction, using the functions for bivariate maps or value-suppressing uncertainty palettes.
Conclusion
This vignette has covered quite a lot of ground, starting from collecting open biodiversity data, and ending at providing interpretable explanations for the output of a species distribution model. There are more functionalities in the package that are not documented here, and notably the integration with other Julia packages.
If there are elements in this tutorial, or in the rest of the documentation, that are not clear, feel free to reach out using the GitHub discussions!
Related documentation
SDeMo.crossvalidate Function
crossvalidate(sdm, folds; thr = nothing, kwargs...)Performs cross-validation on a model, given a vector of tuples representing the data splits. The threshold can be fixed through the thr keyword arguments. All other keywords are passed to the train! method.
This method returns two vectors of ConfusionMatrix, with the confusion matrix for each set of validation data first, and the confusion matrix for the training data second.
crossvalidate(sdm::T, args...; kwargs...) where {T <: AbstractSDM}Performs cross-validation using 10-fold validation as a default. Called when crossvalidate is used without a folds second argument.
SDeMo.outofbag Function
outofbag(ensemble::Bagging; kwargs...)This method returns the confusion matrix associated to the out of bag error, wherein the succes in predicting instance i is calculated on the basis of all models that have not been trained on i. The consensus of the different models is a simple majority rule.
The additional keywords arguments are passed to predict.
SDeMo.explainmodel Function
explainmodel(ModelExplanation, model, variable::Tuple{Int,Int}, n::Int; kwargs...)Generates an n by n grid to measure the surface representing the joint explanation for two variables. ModelExplanation is any type that can be used as the first argument for explainmodel. Other kwargs... are passed to explainmodel.
explainmodel(ModelExplanation, model, variable::Tuple{Int,Int}, n::Int; kwargs...)Generates a grid to measure the surface representing the joint explanation for two variables, where the entries of the grid are the sorted unique values for the pair of variables to consider. ModelExplanation is any type that can be used as the first argument for explainmodel. Other kwargs... are passed to explainmodel.
explainmodel(ModelExplanation, model, variable::Int, n::Int; kwargs...)Generates n equally spaced values over the range of values for the variable of interest, on which the requested explanation will be measured. ModelExplanation is any type that can be used as the first argument for explainmodel. Other kwargs... are passed to explainmodel.
explainmodel(ModelExplanation, model, variable::Int, n::Int; kwargs...)Sorts the unique values for the variable of interest, on which the requested explanation will be measured. ModelExplanation is any type that can be used as the first argument for explainmodel. Other kwargs... are passed to explainmodel.
explainmodel(PartialResponse, model, variable, x; inflated=false, kwargs...)Returns the partial response of the model for a given variable, which is evaluated at all values given by x. The partial response is the prediction made using the given value of variable, when all other variables take the average value. When inflated is true, the response is measured against a randomly sampled value in the entire range of possible values for the other variables.
The result is returned as a tuple x, response.
All other kwargs... are passed to predict.
explainmodel(PartialResponse, model, variable, x, y; inflated=false, kwargs...)Returns the partial response of the model for a given pair of variable given as a tuple, which are evaluated at all values given by vectors x and y.
The result is returned as a tuple x, y, response.
All other kwargs... are passed to predict.
explainmodel(CeterisParibus, model, variable, index, x; kwargs...)Returns the ceteris paribus explanation for a given instance (given as an index: the n-th instance of the model) and a given variable, whose values are evaluated at all positions in the vector x.
Additional kwargs... are passed to predict.
This function returns a tuple x, response.
explainmodel(CeterisParibus, model, variable, index, x, y; kwargs...)Returns the ceteris paribus explanation for a given instance (given as an index: the n-th instance of the model) and a given variable pair, whose values are evaluated at all positions in the vectors x and `y.
Additional kwargs... are passed to predict.
This function returns a tuple x, y, response, where the response is a matrix.
explainmodel(CeterisParibus, model, variable, index, n; kwargs...)Returns the ceteris paribus explanation for a given instance (given as an index: the n-th instance of the model) and a given variable, whose values are evaluated at n equally spaced positions across the range of the variable in the training data.
Additional kwargs... are passed to predict.
This function returns a tuple x, response.
explainmodel(CeterisParibus, model, variable, index; kwargs...)Returns the ceteris paribus explanation for a given instance (given as an index: the n-th instance of the model) and a given variable, whose values are evaluated at all the (sorted) unique values of the variable in the training data.
Additional kwargs... are passed to predict.
This function returns a tuple x, response.
explainmodel(CeterisParibus, model, variable::Tuple{Int,Int}, index, n; kwargs...)Returns the ceteris paribus explanation for a given instance (given as an index: the n-th instance of the model) and a given variable pair, whose values are evaluated at n evenly spaced position across the range of both variables.
Additional kwargs... are passed to predict.
This function returns a tuple x, y, response, where the response is a matrix.
explainmodel(CeterisParibus, model, variable::Tuple{Int,Int}, index; kwargs...)Returns the ceteris paribus explanation for a given instance (given as an index: the n-th instance of the model) and a given variable pair, whose values are evaluated at the (sorted) unique pairs of both variables.
Additional kwargs... are passed to predict.
This function returns a tuple x, y, response, where the response is a matrix.
explainmodel(PartialDependence, model, variable, x; kwargs...)Returns the partial dependence curve of the model for a given variable, which is evaluated at all values given by x.
The result is returned as a tuple x, response.
All other kwargs... are passed to predict.
explainmodel(PartialDependence, model, variable::Tuple{Int, Int}, x, y; kwargs...)Returns the partial dependence curve of the model for a given pair of variables, which is evaluated at all values given by x and y.
The result is returned as a tuple x, y, response.
All other kwargs... are passed to predict.
explainmodel(ShapleyMC, model, variable::Int; kwargs...)Returns the Shapley values (approximation) for the given variable. All other arguments are passed to predict.
The result is returned as a tuple feature, explanation.
explainmodel(ShapleyMC, model, variable::Int, index::Int; kwargs...)Returns the Shapley values (approximation) for the given variable for the given index observation. All other arguments are passed to predict.
The result is returned as a tuple feature, explanation.
explainmodel(ShapleyMC, model, variable::Int, X::Matrix; kwargs...)Returns the Shapley values (approximation) for the given feature matrix X.
explainmodel(ModelExplanation, model, variable, layers; kwargs...)Returms a layer with the explanation of the effect of a given variable, provided by either PartialDependence, PartialResponse, or ShapleyMC.
Note that PartialDependence may take longer to run, as it will run the CeterisParibus explanations for all the training instances at every point in the raster.
All other kwargs... are passed to explainmodel.
explainmodel(ModelExplanation, model, variable, layers; kwargs...)Returms a vector of layers with the explanations for the effect of all variables, provided by either PartialDependence, PartialResponse, or ShapleyMC.
Note that PartialDependence may take longer to run, as it will run the CeterisParibus explanations for all the training instances at every point in the raster.
All other kwargs... are passed to explainmodel.
SDeMo.featureimportance Function
featureimportance(PermutationImportance, model, variable; kwargs...)Returns the feature importance of a variable for a model over the entire model training data. Note that to avoid overfitting, it is better to use a different testing set to evaluate the importance of features.
featureimportance(PermutationImportance, model, variable, X, y; kwargs...)Returns the feature importance of a variable for a model by predicting over features X with a ground truth vector y .
The samples keyword (defaults to 50) is the number of samples to use to evaluate the average importance.
When the ratio keyword is true, the importance is evaluated as the permuted performance divided by the original performance (values lower than 1 indicate that the variable is increasingly important). When it is false, the performance is measured as a difference (values above 0 indicate that the variable is increasingly important).
The optimality keyword takes a function (defaults to mcc) that operates on a a vector of predictions and a vector of correct labels. The interpretation of feature importance assumes that higher values of this measure correspond to a better model.
The other kwargs... are passed to predict. For example, threshold=false, optimality=crossentropyloss is the way to measure the importance of variables based on their effect on cross-entropy loss.
featureimportance(PartialDependence, model, variable; kwargs...)Measures the feature importance of a variable by measuring the distance of all points in the partial dependence curve to all other points in the curve, and then averaging it. Low values indicates a flat PDP (feature is less important). This method can be applied on a fully trained model as it focuses on the response, not the individual predictions.
featureimportance(PermutationImportance, model, variable, training, testing; kwargs...)Calculates the feature importance of a model by re-training a copy of the model using the training instances, then evaluating the feature importance on the testing instances.
Other kwargs... are passed to featureimportance.
featureimportance(ShapleyMC, model, variable; kwargs...)Returns the importance of a feature according to Shapley values; this is the average of the absolute values of this feature on the average prediction (and therefore, this is conceptually very close to the PartialDependence measure of feature importance).
Other kwargs... are passed to predict.
featureimportance(ShapleyMC, model, variable, X::Matrix; kwargs...)Returns the importance of a feature according to Shapley values; this is the average of the absolute values of this feature on the average prediction (and therefore, this is conceptually very close to the PartialDependence measure of feature importance). The Shapley values are measured on the matrix of instances X.
Other kwargs... are passed to predict.
PseudoAbsences.pseudoabsencemask Function
pseudoabsencemask(::Type{RandomSelection}, presences::SDMLayer{Bool})Generates a mask for pseudo-absences using the random selection method. Candidate cells for the pseudo-absence mask are (i) within the bounding box of the layer (use SurfaceRangeEnvelope to use the presences bounding box), and (ii) valued in the layer.
pseudoabsencemask(::Type{SurfaceRangeEnvelope}, presences::SDMLayer{Bool})Generates a mask from which pseudo-absences can be drawn, by picking cells that are (i) within the bounding box of occurrences, (ii) valued in the layer, and (iii) not already occupied by an occurrence
sourcepseudoabsencemask( ::Type{DistanceToEvent}, presences::SDMLayer{Bool}; f = minimum, )Generates a mask for pseudo-absences using the distance to event method. Candidate cells are weighted according to their distance to a known observation, with far away places being more likely. Depending on the distribution of distances, it may be a very good idea to flatten this layer using log or an exponent. The f function is used to determine which distance is reported (minimum by default, but can be any other relevant function).
pseudoabsencemask(::Type{WithinRadius}, presences::SDMLayer{Bool}; distance::Number = 100.0, )Generates a mask for pseudo-absences where pseudo-absences can be within a distance (in kilometers) of the original observation. Internally, this uses DistanceToEvent. All other keyword arguments are passed to pseudoabsencemask.
pseudoabsencemask(::Type{WithoutRadius}, presences::SDMLayer{Bool}; distance::Number = 100.0, )Generates a mask for pseudo-absences where pseudo-absences can be outside of a distance (in kilometers) of the original observation. Internally, this uses DistanceToEvent. All other keyword arguments are passed to pseudoabsencemask.
pseudoabsencemask(::Type{BetweenRadius}, presences::SDMLayer{Bool}; closer::Number = 10.0, further::Number=100.0)Generates a mask for pseudo-absences where pseudo-absences can be in a band given by two distances (in kilometers) of the original observation. Internally, this uses DistanceToEvent. All other keyword arguments are passed to pseudoabsencemask.
pseudoabsencemask( ::Type{DegreesToEvent}, presences::SDMLayer{Bool}; f = minimum, absolute::Bool = false)Generates a mask for pseudo-absences using the distance to event method. Candidate cells are weighted according to their distance to a known observation, with far away places being more likely. The f function is used to determine which distance is reported (minimum by default, but can be any other relevant function). The distance is measured in degrees, and is therefore very sensitive to the projection / position.
When absolute is false (the default), the distance is measured as the absolute difference in coordinates instead of using the Euclidean distance.
pseudoabsencemask(::Type{WithinDegrees}, presences::SDMLayer{Bool}; distance::Number = 3.0, )Generates a mask for pseudo-absences where pseudo-absences can be within a distance (in degrees) of the original observation. Internally, this uses DegreesToEvent. All other keyword arguments are passed to pseudoabsencemask.
pseudoabsencemask(::Type{WithoutDegrees}, presences::SDMLayer{Bool}; distance::Number = 3.0, )Generates a mask for pseudo-absences where pseudo-absences can be outside of a distance (in degrees) of the original observation. Internally, this uses DegreesToEvent. All other keyword arguments are passed to pseudoabsencemask.
pseudoabsencemask(::Type{BetweenDegrees}, presences::SDMLayer{Bool}; closer::Number = 1.0, further::Number=3.0)Generates a mask for pseudo-absences where pseudo-absences can be in a band given by two distances (in degrees) of the original observation. Internally, this uses DegreesToEvent. All other keyword arguments are passed to pseudoabsencemask.
PseudoAbsences.backgroundpoints Function
backgroundpoints(layer::T, n::Int; kwargs...) where {T <: SimpleSDMLayer}Generates background points based on a layer that gives the weight of each cell in the final sample. The additional keywords arguments are passed to StatsBase.sample, which is used internally. This includes the replace keyword to determine whether sampling should use replacement.
GBIF.taxon Function
Get information about a taxon at any level
taxon(name::String)This function will look for a taxon by its (scientific) name in the GBIF reference taxonomy.
Optional arguments are
rank::Union{Symbol,Nothing}=:SPECIES– the rank of the taxon you want. This is part of a controlled vocabulary, and can only be one of:DOMAIN,:CLASS,:CULTIVAR,:FAMILY,:FORM,:GENUS,:INFORMAL,:ORDER,:PHYLUM,,:SECTION,:SUBCLASS,:VARIETY,:TRIBE,:KINGDOM,:SUBFAMILY,:SUBFORM,:SUBGENUS,:SUBKINGDOM,:SUBORDER,:SUBPHYLUM,:SUBSECTION,:SUBSPECIES,:SUBTRIBE,:SUBVARIETY,:SUPERCLASS,:SUPERFAMILY,:SUPERORDER, and:SPECIESstrict::Bool=true– whether the match should be strict, or fuzzy
Finally, one can also specify other levels of the taxonomy, using kingdom, phylum, class, order, family, and genus, all of which can either be String or Nothing.
If a match is found, the result will be given as a GBIFTaxon. If not, this function will return nothing and give a warning.
Get information about a taxon at any level using taxonID
taxon(id::Int)This function will look for a taxon by its taxonID in the GBIF reference taxonomy.
sourceSimpleSDMLayers.mosaic Function
mosaic(f, stack::Vector{<:SDMLayer})Returns a layer that is the application of f to the values at each cell in the array of layers given as the second argument.
SimpleSDMLayers.mosaic(f, layer, polygons)Overload of the mosaic function where the layer is split according to the polygons given as the last argument, and then the function f is applied to all the zones defined this way. The f function can take both positional and keyword arguments.
SimpleSDMLayers.mosaic(f, layer, fc, key, args...; kwargs...)Overload of the mosaic function where the layer is split according to the polygons given as the last argument, and then the function f is applied to all the zones defined this way. The f function can take both positional and keyword arguments.
SDeMo.hyperparameters Function
hyperparameters(::Type{<:HasHyperParams}) = nothingReturns the hyper-parameters for a type of classifier or transformer
sourcehyperparameters(::HasHyperParams, ::Symbol)Returns the value for an hyper-parameter
sourcehyperparameters(::HasHyperParams)Returns the hyper-parameters for a classifier or a transformer
sourceSDeMo.hyperparameters! Function
hyperparameters!(tr::HasHyperParams, hp::Symbol, val)Sets the hyper-parameters for a transformer or a classifier
sourceSimpleSDMLayers.SDMLayer Type
SDMLayer{T}Defines a layer of geospatial information.
The type has two data fields:
grid: a
Matrixof typeTindices: a
BitMatrixto see which positions are valued
Each row in the grid field represents a slice of the raster of equal northing, i.e. the information is laid out in the matrix as it would be represented on a map once displayed. Similarly, columns have the same easting.
The geospatial information is represented by three positional fields:
x and y: two tuples, indicating the coordinates of the corners alongside the x and y dimensions (e.g. easting/northing) - the default values are
(-180., 180.)and(-90., 90.), which represents the entire surface of the globe in WGS84crs: any
Stringrepresentation of the CRS which can be handled byProj.jl- the default is EPSG:4326
Note that we recommend that the SRS be represented as WKT, then as the "EPSG:xxx" format, and then as a a PROJ4 string, by order of decreasing preference. An ArchGDAL representation is always accessible through the projection method. Note that the functions will, internally, convert the projection to different formats in order to achieve specific tasks.
SDeMo.SDM Type
SDMThis type specifies a full model, which is composed of a transformer (which applies a transformation on the data), a classifier (which returns a quantitative score), a threshold (above which the score corresponds to the prediction of a presence).
In addition, the SDM carries with it the training features and labels, as well as a vector of indices indicating which variables are actually used by the model.
The coordinates for each observation that is used to train the model are given in the coordinates field, and as with OccurrencesInterface, they must be given as longitude,latitude. If there are no known coordinates for the observations, this field must be an empty vector of the correct type. As of now, there is no plan to support datasets that only have some coordinates known.
SDeMo.Bagging Type
BaggingA bagged (bootstrap aggregated) model is the same template model repeated an arbitrary number of times, with each replicated model having access to a bootstrapped sample of the data. These models are represented by three fields:
modelis the base modelbagsis a vector of tuples with the in and out-of sample instancesmodelsis an array of replicated models